Command Palette
Search for a command to run...
Based on 25,000 Clinical Data Points, Stanford University Released the First Native 3D Abdominal CT Visual Language Model, Merlin, Which Leads Across 752 tasks.

Computed tomography (CT) is a commonly used imaging technique in clinical diagnosis and treatment, and it is widely used in the diagnosis of diseases in multiple parts of the body. Statistics show that approximately 300 million CT scans are performed globally each year, with abdominal CT scans accounting for about one-quarter. As medical diagnosis and treatment become increasingly reliant on imaging technology, the demand for imaging diagnostics continues to grow. However, it typically takes a radiologist 20 minutes to interpret a single abdominal CT image, and the diagnostic efficiency is struggling to keep pace with the rapidly increasing clinical demand. Even more serious is the severe shortage of radiologists; predictive data shows that…By 2036, some regions will face a shortage of more than 19,000 radiologists, highlighting the growing imbalance between supply and demand in the industry.
Machine learning, with its sophisticated data processing and high-throughput analysis capabilities, can rapidly extract features and intelligently identify massive amounts of medical images, effectively addressing the pain points of traditional manual image interpretation, such as low efficiency and insufficient manpower. In particular, Vision-Language Models (VLMs), driven by Contrastive Language-Image Pretraining (CLIP) pretraining technology, enable text representations and visual representations to be aligned in a shared embedding space, thereby supporting the supervision of visual models using natural language.As a foundational model, this type of model can not only achieve zero-shot learning, but also, after being combined with a large language model and trained with clinical data, can be quickly adapted to the analysis of radiological images and reports.
Beyond theoretical and technological advancements, current VLM-based methods are demonstrating immense application potential in the field of radiology, with models such as BiomedCLIP, LLaVA-Rad, and Med-PaLMM being successfully implemented. However, technological progress and model implementation do not equate to mature application. VLMs still face numerous key challenges in practical applications, hindering their widespread adoption and reliable use in clinical settings.
first,Existing methods mostly focus on two-dimensional images such as X-ray films, making it difficult to efficiently process three-dimensional images such as abdominal CT scans. The method of analyzing the full volume through slice aggregation is extremely inefficient.Secondly,Currently, there is no publicly available abdominal CT dataset for training and evaluation of VLMs. Private models have not fully integrated multimodal clinical data such as diagnostic coding and radiology reports, and there is a lack of a unified three-dimensional abdominal CT task benchmark, resulting in a significant gap in the training and evaluation system of related basic models.
In view of the above challenges,A research team from Stanford University has proposed Merlin, the first native 3D visual language model for abdominal CT scans, along with a dataset containing 25,494 paired abdominal CT scans and radiology reports. Merlin was trained on a single NVIDIA A6000 GPU using structured and unstructured data from real-world hospitals, including paired CT scans, Electronic Health Record (EHR) diagnostic codes, and radiology reports. The research team performed internal validation on 5,137 CT scans and external validation on 44,098 CT scans and two publicly available datasets focusing on abdominal CT scans (VerSe and TotalSegmentator). The validation results show that Merlin comprehensively outperforms specific benchmark models on benchmark tasks.
The related research findings, titled "Merlin: a computed tomography vision–language foundation model and dataset," were published in Nature.
Research highlights:
* This study proposes Merlin, the first native 3D visual language model specifically for abdominal CT scans, overcoming the limitation of previous models that only focused on 2D images.
* The study released a large-scale dataset containing 25,494 paired abdominal CT scans and radiology reports, filling a gap in the dataset field.
* This research innovatively integrates structured EHR data and unstructured radiology reports as supervisory signals, and proposes a multi-stage pre-training framework that combines multi-task learning and phased training.

Paper address:
https://www.nature.com/articles/s41586-026-10181-8
Follow our official WeChat account and reply "Merlin" in the background to get the full PDF.
Filling the data gap for training and evaluation of VLMs
To fill the gap in the lack of publicly available abdominal CT datasets for training and evaluating 3D VLMs, the research team used a large amount of compliant data from real medical centers.Ultimately, a high-quality clinical dataset containing 18,321 patients was released, covering paired CT scans, unstructured radiology reports, and structured EHRs.in:
* CT scan data:
The data were derived from whole abdominal CT scans, each containing multiple sequences. The sequence with the most axial slices was selected to maximize information content. This process yielded 10,628,509 two-dimensional images from 25,528 CT scans.
* Radiological report:
The study compiled the radiology reports corresponding to each CT scan. These reports consist of multiple parts, the most crucial being "findings" and "impressions." The former includes detailed observations of each organ system, while the latter summarizes key clinical findings. Notably, based on the granularity of the provided information and the validity of previous work, the training only utilized the "findings" section, totaling 10,051,571 tokens.
* EHR:
The data was used to train the model using diagnostic information in the form of International Classification of Diseases (ICD) codes, which were associated with the corresponding patients' CT scan records. The dataset contains a total of 954,013 ICD9 codes, involving 5,686 unique codes; and 2,041,280 ICD10 codes, involving 10,867 unique codes.
In terms of data partitioning, the pre-training dataset was divided into three sub-databases: 60% (15,331 CT scans), 20% (5,060 CT scans), and 20% (5,137 CT scans), used for training, validation, and testing, respectively. As a precaution, multiple CT scans of a single patient were not included in the same sub-database.
also,The experiment also used 44,098 data points from three independent institutions for external validation, all of which were used for testing.The details are as follows:
* External dataset 1: Contains 6,997 abdominal CT scans
* External dataset 2: Contains 25,986 abdominal CT scans
* External dataset 3: Contains 4,872 abdominal CT scans and 6,243 chest CT scans.
The other two dedicated public datasets for abdominal CT scans are VerSe and TotalSegmentator. The VerSe dataset contains 160 CT scans, while the TotalSegmentator dataset contains 401 CT scans. Of these, 34 selected scans were used for pre-training and testing of multi-task, multi-disease prediction, and the remaining 367 scans were divided into 80% (293 scans) and 20% (74 scans) for training and validation, respectively.
Multi-task learning and phased training strategies, differentiated solutions ensure Merlin's high efficiency.
In terms of model architecture,Merlin achieves image-text alignment by employing a dual encoder architecture consisting of an image encoder and a text encoder.The image encoder uses I3D ResNet152, which reuses the weights of the two-dimensional pre-trained model through "inflation" and copies them to the third dimension of the three-dimensional convolutional kernel. The encoder used in this paper is Clinical Longformer, which has a longer text capability than other biomedical pre-trained models and general CLIP encoders, supporting 4,096 long contexts and adapting to the needs of long text reports.
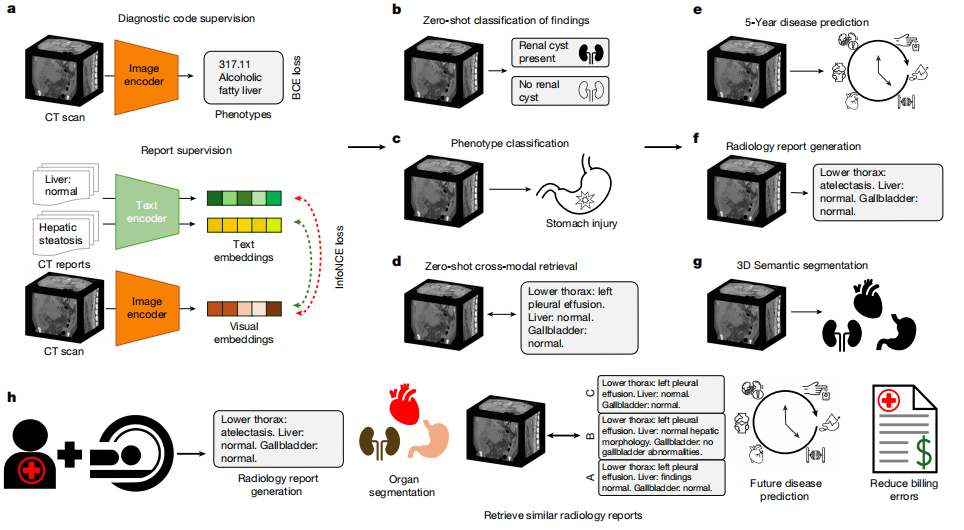
For model training, Merlin uses two loss functions to handle phenotypic classification and radiological reports respectively:The binary cross-entropy loss function was used for phenotypic classification; the InfoNCE loss function was used for radiological report contrast learning.The embedding dimension for both images and text was uniformly set to 512, consistent with the embedding dimension used in the ViT-Base model in the OpenCLIP experiments. Subsequently, gradient checkpointing was enabled for both the visual encoder and the text encoder in the training strategy, and FP16 mixed-precision training was employed.
The optimizer used was AdamW with an initial learning rate of 1 x 10⁻⁵ and β = (0.9, 0.999). A cosine learning rate scheduler was employed, setting the number of training epochs at which the learning rate decayed to 0 to 300. The hardware consisted of a single 48GB A6000 GPU, with a maximum batch size of 18.
In addition to training using EHR phenotypes and radiological reports in a multi-task manner,The study also considered a phased training program.Specifically, the Merlin image encoder is first trained using EHR diagnostic codes in the first stage; then, it is trained comparatively using radiological reports in the second stage. To prevent the EHR information learned in the first stage from being forgotten, the phenotypic loss function is incorporated with lower weights in the second stage training.
The first stage uses the AdamW optimizer with an initial learning rate of 1 x 10⁻⁴, β = (0.9, 0.999), an exponential learning rate scheduler with γ = 0.99, and a single A6000 GPU with a batch size of 22. The hyperparameters used in the second stage are the same as those used in multi-task training.
In summary, multi-task learning and phased training achieve differentiated designs for the two strategies, and the research team made anti-forgetting improvements for phased training. This differentiated training strategy is the core design that ensures Merlin's efficiency and rigor, and it was further validated in subsequent ablation experiments.
A comprehensive evaluation of 752 task categories shows that Merlin outperforms all others.
In the experimental process, the research team conducted internal validation based on 5,137 CT scans and external validation based on 44,098 CT scans and two publicly available datasets (VerSe and TotalSegmentator) focusing on abdominal CT scans.There are a total of 6 major categories of assessment tasks, covering 752 specific sub-tasks.The major task categories include zero-shot classification (31 sub-tasks), phenotypic classification (692 sub-tasks), zero-shot cross-modal retrieval (23 sub-tasks), 5-year disease prediction (6 sub-tasks), radiology report generation, and 3D segmentation.
In the zero-shot classification of findings task, 30 abdominal CT scans from internal and external clinical data were analyzed.Merlin achieved an F1 score of 0.741 on the internal validation dataset (confidence interval of 95%, 0.727–0.755) and an average F1 score of 0.647 on the external validation dataset (confidence interval of 95%, 0.607–0.678).These scores were significantly higher than the 2D OpenCLIP model using k=1 pooling and the finely tuned 2D BioMedCLIP model using average pooling (P < 0.001). See the figure below:
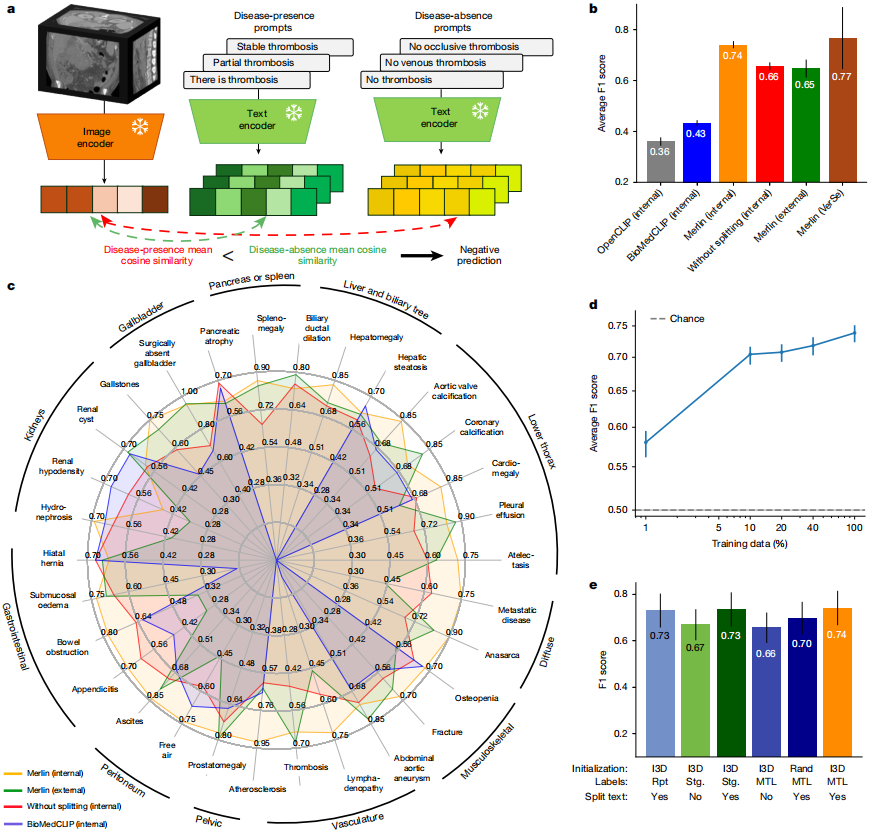
From a qualitative perspective,Merlin maintains high performance on external datasets for diseases with significant features, such as pleural effusion and ascites.However, its performance slightly decreases when dealing with the detection of fine-grained features, such as appendicitis and lymphadenopathy. In addition, without radiology report segmentation, Merlin achieved an average F1 score of 0.656 (confidence interval of 95%) on the external evaluation dataset.
In the ablation experiment comparisonThe Merlin model initialized with an dilated 3D network exhibits the best performance.The F1 score was 0.741 (confidence interval of 95%, 0.727–0.755); when segmenting radiology reports, the model score combining EHR and radiology reports was 0.735 (confidence interval of 95%, 0.719–0.748); the scheme using only radiology reports and implementing report segmentation ranked third, with an F1 score of 0.730 (confidence interval of 95%, 0.714–0.744). Whether or not radiology reports were segmented had the most significant impact on model performance; without report segmentation, the F1 score of the Merlin model decreased by an average of 7.9 points (P < 0.01).
It is also worth mentioning thatZero-shot Merlin outperforms all supervised baselines in supervised experiments on both 10% and 100% training data.When using 100% training data, the F1 score improved by 29%, while when using 10% training data, it improved by a staggering 45%. Experiments demonstrate that, with 100% training data, zero-shot Merlin significantly outperforms supervised Merlin, improving the F1 score by 16%.
In the phenotype classification task, Merlin's performance in predicting 692 clinical phenotypes defined by PheWAS was evaluated, achieving a macro-mean area under the receiver operating characteristic (AUROC) of 0.812 (95% confidence interval, 0.808–0.816). A total of 258 phenotypes had AUROC values exceeding 0.85, and 102 phenotypes had AUROC values exceeding 0.9. (See figure below.)
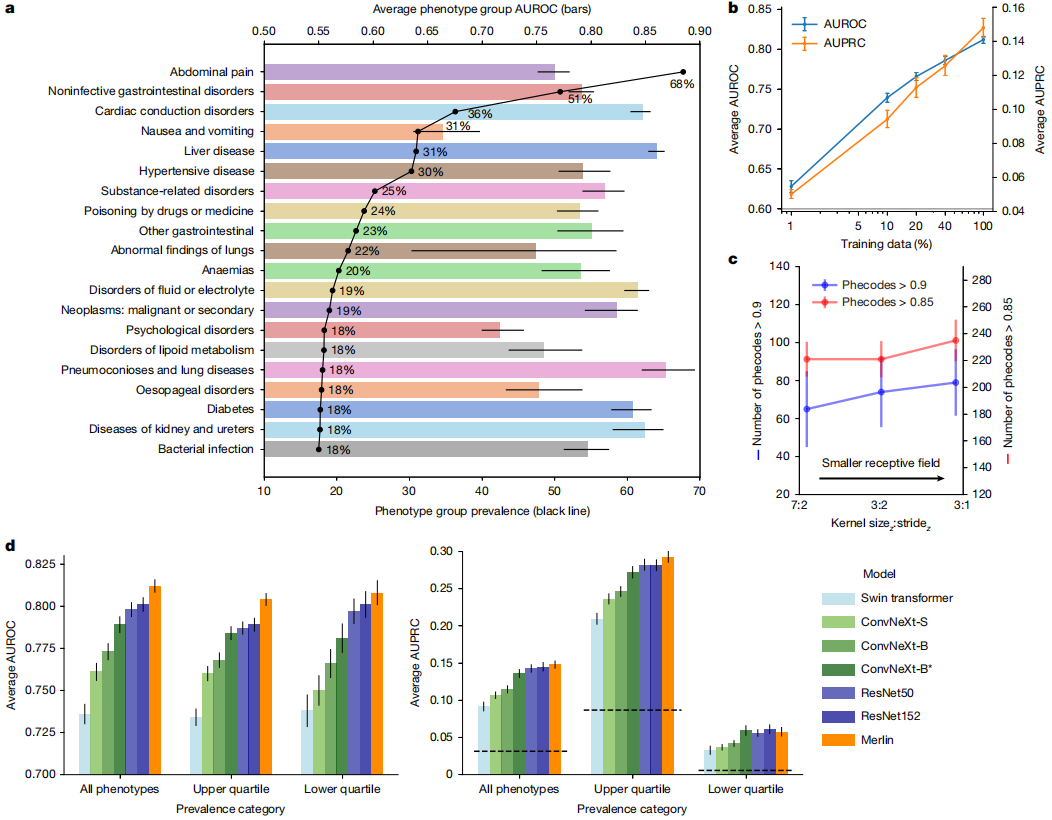
When analyzing the top 20 most common phenotypes with the highest incidence rates in internal testing,Merlin excels in detecting diseases of multiple organ systems, including the liver, kidneys, ureters, and gastrointestinal tract.
In the zero-shot cross-model retrieval task, the first step is a retrieval task based on "image discovery" with 64 cases.Merlin exhibits significant advantages over OpenCLIP and BioMedCLIP.This is thanks to the Clinical Longformer text encoder used by Merlin, while OpenCLIP and BioMedCLIP allow maximum token lengths of 77 and 256, respectively. Conversely, Merlin's excellent performance was also replicated in the "discovery-image" retrieval task based on 64 cases. See the figure below:
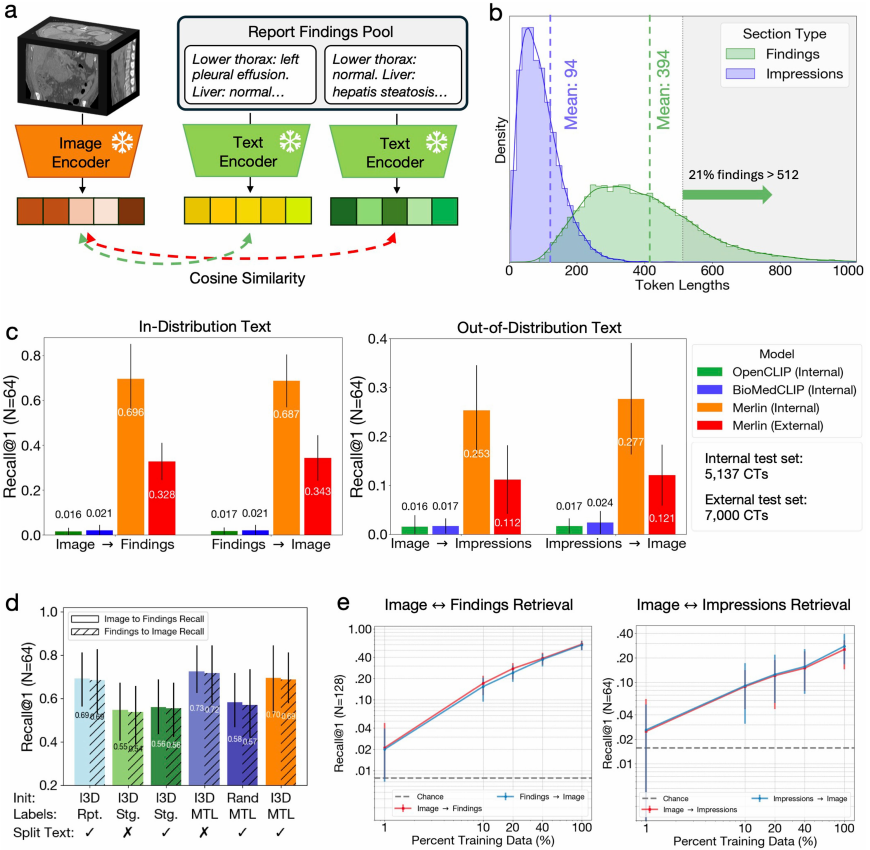
More important evidence is that Merlin, even when using only the objectively described "findings" in the report for visual-language alignment training,Even when dealing with highly generalized report "impressions," it still demonstrates a high degree of cross-domain generalization ability.The results were then verified again in a reverse engineering task. Furthermore, although Merlin's retrieval performance on the external test dataset decreased compared to the internal test dataset, it was still 5-7 times better than other external baselines.
In the Multi-disease 5-year prediction task, the experiment assessed Merlin's prediction of the risk of healthy patients developing multiple major chronic diseases over the next five years, including chronic kidney disease, osteoporosis, cardiovascular disease, ischemic heart disease, hypertension, and diabetes.
After fine-tuning Merlin and using the 100% downstream label, its AUROC value for predicting disease incidence within five years reached 0.757 (confidence interval of 95%, 0.743-0.772).This performance is 71 TP3T higher than the ImageNet pre-trained (I3D) model that only uses images.Even using only 10% labels, Merlin's AUROC for predicting disease incidence within five years still reaches 0.708 (confidence interval of 95%, 0.692-0.723), outperforming the ImageNet pre-trained model's 4.4%. See the figure below:
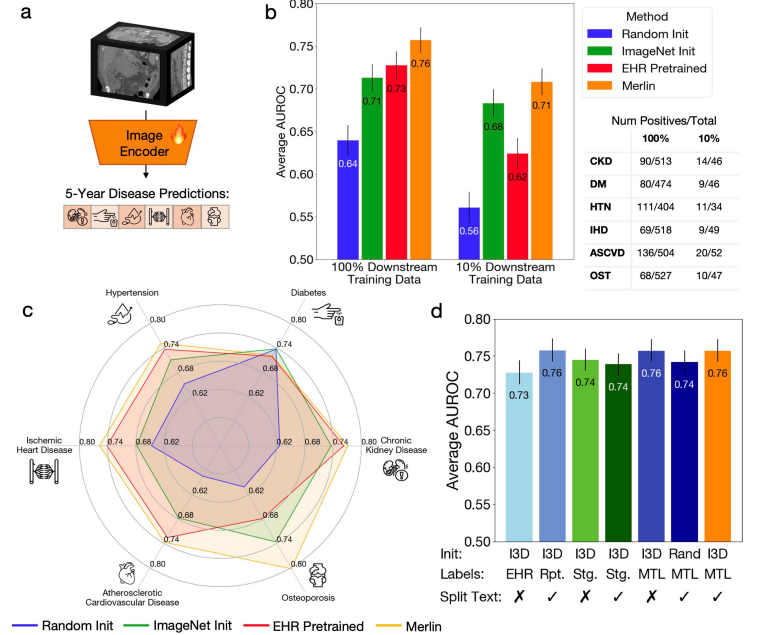
in addition,Even using only 1/10 of the training data, Merlin's prediction performance is comparable to that of an ImageNet pre-trained model trained on 100% of data.This greatly demonstrates Merlin's zero-shot capability and powerful transferability.
In the radiology report generation task, compared to the baseline model RadFM, in tests based on quantitative metrics such as RadGraph-F1, BERT Score, ROUGE-2, and BLEU,Merlin outperforms the former in all aspects of anatomical logical structure and complete reporting results.
In terms of quality, Merlin generates excellent reports with highly accurate diagnosis, localization, and description of symptoms. However, Merlin occasionally makes conservative judgments, such as underreporting issues found in both manually generated and CT reports. This is due to early demonstrations of radiology reports generated from CT scans, and will be further improved as report quality increases.
In the 3D semantic segmentation task, Merlin outperforms the nnUNet framework by 4.71 TP3T in macro-average Dice score when using only 101 TP3T of training data; when using 1001 TP3T of training data, the nnUNet framework performs slightly better than Merlin's initial model, but the difference in Dice score is only 0.006.
On 20 organs in the test set, Merlin achieved higher Dice scores than the nnUNet framework on 12 organs when trained with 10% data, with an improvement of up to 41% in prostate segmentation.
In addition, in external validation trials, the research team evaluated Merlin on a total of 44,098 external CT scans using a dataset of over 100,000 external CT scans.It exhibits stable and accurate performance across different sites and anatomical locations, overcoming the distribution offset between the training dataset and the external test dataset.Furthermore, its performance consistently outperformed other baseline models, even surpassing specialized chest CT baseline models on chest tasks.
Visual language models unlock the potential value of large-scale multimodal medical data.
In addition to this study, other achievements in visual language models in medicine are emerging one after another. For example, a research team from Stanford University proposed a multimodal transformer with unified masked modeling (MUSK), which is also a basic model of visual language and aims to integrate large-scale, unlabeled, unpaired image and text data.
Paper Title: A vision–language foundation model for precision oncology
Paper address:
https://www.nature.com/articles/s41586-024-08378-w
The knowledge-enhanced case-based model KEEP, proposed by Shanghai Jiao Tong University and others, addresses the problem that current models primarily rely on data-driven approaches and lack explicit integration of medical knowledge. This model utilizes a comprehensive disease knowledge graph of 11,454 diseases and 139,143 attributes to reorganize millions of pathological image-text pairs into 143,000 semantically structured groups aligned with the disease ontology hierarchy. This knowledge-enhanced pre-training method aligns visual and textual representations in a hierarchical semantic space, thereby achieving a deep understanding of disease relationships and morphological patterns.
Thesis title: Knowledge-enhanced pretraining for vision-language pathology foundation model on cancer diagnosis
Paper address:
https://www.sciencedirect.com/science/article/pii/S1535610826000589
In summary, visual language models, with their cross-modal understanding capabilities, are demonstrating tremendous potential in the fields of medicine and radiology. They can integrate medical images, case texts, and clinical guidelines to achieve intelligent lesion identification, case analysis assistance, and automatic generation of diagnostic reports. This not only provides physicians with efficient auxiliary tools but also offers new insights into disease prediction, accelerating the shift of modern medicine from "experience-driven" to "data-driven."