Command Palette
Search for a command to run...
Based on Clinical Data From 11,647 Cases, a French Team Has Achieved, for the First Time, Accurate Prediction of the Dual Mortality Risk in HCC Liver Transplantation Using Machine learning.
Liver cancer, due to its insidious early stages and rapid progression, has long been known as the "king of cancers." Among them, hepatocellular carcinoma (HCC) is the most common type of liver cancer, accounting for 70%–90% of primary liver cancers. Patients usually need liver transplantation as a radical treatment in the early stages, which is also the last "lifeline" for many HCC patients to grasp at the hope of life.
However, the extreme scarcity of donor organs makes this hope for life all the more precious. Even more challenging is the fact that HCC liver transplant candidates constantly face the dual threat of death from liver failure and tumor progression; these two factors are intertwined and mutually influential, greatly increasing the risk of death during the waiting period. Therefore,Accurately assessing the risk of death during the waiting period for HCC liver transplant candidates is not only key to optimizing the priority of the liver transplant waiting list and achieving fair allocation of scarce donors, but also a core challenge to efficiently save every patient and safeguard the hard-won hope of life.
Previously, traditional risk assessment methods such as Child-Pugh, Albumin-Bilirubin (ALBI), and Model for End-Stage Liver Disease (MELD) were widely used in liver disease risk assessment. However, they showed significant shortcomings when faced with the complex situation of HCC patients: these methods either focused on assessing liver function and the degree of cirrhosis or only focused on predicting tumor progression, failing to address both risks simultaneously. Even with the subsequent development of comprehensive scoring systems such as HALT-HCC and the Mehta Model, which can consider both risks at the same time,Also, due to limitations of linear models, fixed variable weights, and static measurements at a single time point, it is impossible to capture the interactions between influencing factors and the risk changes in the dynamic progression of the disease, making it difficult to achieve accurate individualized risk assessment.
In response to this clinical pain point,A research team from Telecom Sud-Paris and Paris-Saclay University in France has proposed a machine learning framework that integrates ensemble learning (EL) with Schapel Additive exPlanations (SHAP) analysis.This study provides a new approach to assessing mortality risk in HCC liver transplant candidates. Based on clinical data from 11,647 patients, it compared three ensemble models: Random Forest (RF), XGBoost, and LightGBM. Furthermore, by embedding SHAP values into the Uniform Manifold Approximation and Projection (UMAP) low-dimensional space and combining it with K-medoids algorithm for supervised clustering, the study clarified that liver dysfunction and tumor progression are the two core risks of death in HCC patients.
This study specifically fills a gap in previous machine learning models for accurately assessing HCC liver transplant candidates, especially in studies involving dual risks.This study achieves accurate prediction and clinical interpretability of mortality during the 3-month waiting period for HCC liver transplant candidates, providing a new tool for clinical decision-making and risk stratification in HCC patients undergoing liver transplantation.
The findings, titled "Explainable Mortality Prediction for Liver Transplant Candidates with Hepatocellular Carcinoma: A Supervised Clustering Approach," were published in Health Data Science.
Research highlights:
* This study is the first comprehensive study to use machine learning models to deeply analyze the mortality risk of HCC liver transplant candidates on the waiting list.
* By using SHAP + UMAP + K-medoids, seven clinically explainable risk subgroups were stratified to identify the core drivers of dual risk.
* The novel risk score ELM-HCC, constructed based on SHAP screening of 8 key variables, demonstrates significantly superior predictive accuracy compared to traditional scores.
* This study is the first to incorporate key dynamic variables (such as AFP_DIFF) into the risk assessment of HCC liver transplant candidates, clarifying their role as a key predictor of mortality during the waiting period for HCC patients.
Paper address:
https://spj.science.org/doi/10.34133/hds.0295
Follow our official WeChat account and reply "liver transplant" in the background to get the complete PDF.
View more cutting-edge AI papers:
Dataset: Large Sample Strategy + Dynamic Variable Introduction
To reduce confounding factors,The study employed a large-sample strategy based on public database data.
Specifically, the study data came from the Standard Transplant Analysis and Research (STAR) files of the Organ Procurement and Transplantation Network (OPTN) and the United Network for Organ Sharing (UNOS), covering adult HCC patients who did not undergo multiple organ transplantation and were registered between February 27, 2002 and September 30, 2023.
This study aimed to predict mortality during the 3-month waiting period for liver transplantation in HCC patients. Therefore, the research team divided the study population into two groups for analysis.Patients who have been on the waiting list for more than three months are referred to as "on the waiting list"; patients who die on the waiting list within three months or whose condition worsens and makes them unable to receive a transplant are referred to as "waitlist mortality".final,The total study cohort included 11,647 patients.Of these, 11,199 patients were on the waiting list and 448 patients were on the waiting list for mortality. The data included clinical, laboratory, and disease-related multidimensional variables.
In the data preprocessing stage, to capture the dynamic characteristics of patients' health status, the research team calculated the continuous measurement difference (DIFF) of six key laboratory variables involved in traditional scoring, including serum sodium, creatinine, albumin, bilirubin, alpha-fetoprotein (AFP), and the International Normalized Ratio (INR), in order to capture the dynamic trajectory of changes in patients' health status.This increases the total number of features to 31 (25 original static variables + 6 newly added dynamic variables).
For handling missing values, numerical variables (missing rate < 7%) were imputed using the class mean; categorical variables (missing rate < 0.1%) had their observation records containing missing values directly deleted.
Model Architecture: End-to-End Integrated Process + Comparison of Multiple Ensemble Learning Models
To ensure the reliable accuracy and interpretability of the prediction of mortality during the 3-month waiting period for HCC liver transplant candidates,The research team constructed an end-to-end integrated process that combines ensemble learning, SHAP interpretability analysis, UMAP dimensionality reduction, and K-Medoids supervised clustering.As shown in the following figure:
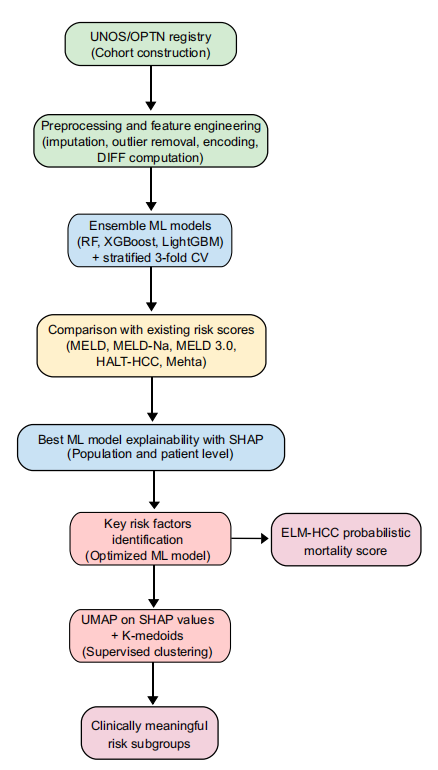
First, the core model employs an ensemble learning tree model.These types of models are particularly effective for handling tabular and heterogeneous data. To further compare the performance of these models, the study used three basic ensemble learning models: Random Forest, XGBoost, and LightGBM. The experiments were conducted in two training scenarios: the first used only 25 original static variables; the second used 31 combined static and dynamic variables, including dynamic variables.
Secondly, the purpose of interpretability is to provide a scientific and reasonable interpretation of the predicted results, thereby enhancing the basis for clinical decision-making.To this end, the research team incorporated SHAP interpretability analysis into the framework to identify key risk factors and reveal model predictions.
For global interpretation, calculating SHAP values quantifies the contribution of each feature to the model's prediction results, identifying core risk factors for mortality prediction and clarifying the correlation between features and mortality risk. For local interpretation, SHAP summary plots and SHAP force plots can demonstrate the specific impact of individual feature values on prediction results, as well as the distribution of feature contributions for each patient. Furthermore, this step provides a SHAP value feature set for subsequent cluster analysis, replacing the original data and improving the clinical interpretability of the clustering.
Finally, to achieve refined risk stratification for patients, the focus shifted from population-level prediction to subgroup-specific analysis.The research process incorporated UMAP dimensionality reduction and K-Medoids supervised clustering methods.First, the predicted SHAP values are embedded into the UMAP dimensionality-reduced space. Then, the K-Medoids algorithm is used to cluster the SHAP values embedded in the 3D UMAP space to discover potential patient subgroups with different clinical characteristics. This method is called "supervised clustering" because the clustering is based on SHAP values rather than the original data.
The optimal number of clusters was determined by first screening using quantitative indicators such as the Silhouette coefficient and Davies-Bouldin index, and then clinically validating the clustering characteristics through SHAP analysis. Ultimately, the optimal number of clusters was determined to be 7.
Experimental Results: The new model was trained using 8 traditional rating methods as a comparison and the optimal feature set.
Risk score performance comparison
The study compares the performance of the proposed framework with eight traditional risk assessment methods, including ALBI, Child–Pugh, AFP, Hazard associated with LT for HCC (HALT-HCC), Mehta Model, MELD and its two variants MELD-Na and MELD 3.0.
Considering the severe class imbalance in the dataset, the study downsampled the majority group (on the waiting list) to generate 30 subsets of similar size to the minority group (deaths during the waiting period). Three-fold cross-validation was performed on each balanced subset to ensure that all observations from the same patient were assigned to either the training or test set. The optimal hyperparameter configurations for the three ensemble models were then determined using a grid search, as shown in the figure below.
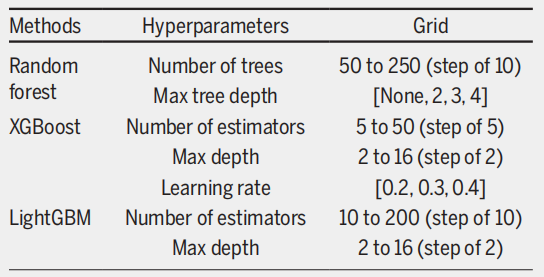
The results show thatIn traditional scoring systems, the Mehta Model performs best with an AUROC of 0.782, followed by the HALT-HCC with an AUROC of 0.763.More importantly, these two models offer a more balanced performance in terms of sensitivity and specificity. While MELD 3.0 outperforms the basic MELD and MELD-Na models, it suffers from an imbalance in sensitivity and specificity.
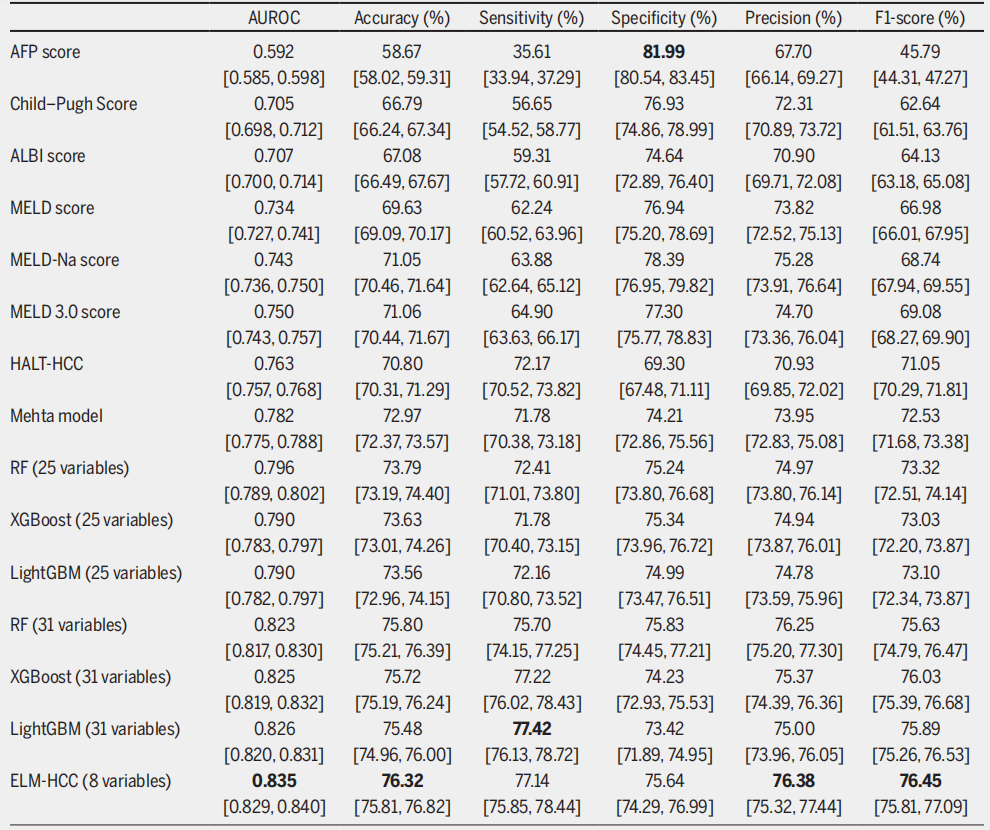
When the experiment was extended to an ensemble learning framework, the accuracy of all models trained on 25 static variables surpassed that of traditional scoring systems. RF performed best, with an AUROC of 0.796, and its sensitivity of 72.41% and specificity of 75.24% were also well-balanced. After introducing 31 combined dynamic and static variables, all ensemble learning models achieved even better performance.LightGBM achieved an AUROC of 0.826 and the highest sensitivity of 77.42%, making it the most effective model for identifying high-risk patients.
Analysis of the ability to identify key risk factors
After the models are trained, the research will use only the most relevant features to evaluate their performance. To this end, the research team used two feature importance evaluation methods, Gain importance and SHAP global importance, to screen key features for the best-performing LightGBM model.
Based on the LightGBM model (the best-performing model), the top 8 features selected using SHAP global importance achieve optimal model performance.With an AUROC of 0.835, a sensitivity of 77.141 TP3T, and a specificity of 75.641 TP3T, it not only outperformed the Gain importance screening results (AUROC of 0.812 with 8 features and reaching its best at 0.828 with 12 features), but also surpassed the performance of LightGBM on a complete set of 31 variables (AUROC of 0.826). Therefore, it was selected by the research team as the optimal feature set.
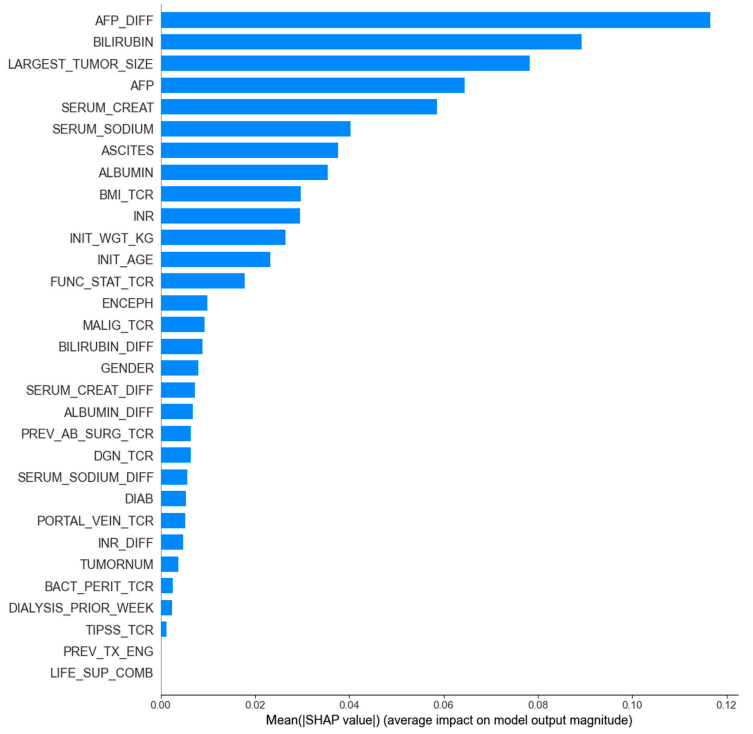
Ultimately, the study identified and constructed a probabilistic mortality score for HCC patients, named ELM-HCC, based on the LightGBM model trained with the optimal feature set. It is worth mentioning that...LightGBM outperformed AUROC on the simplified variable set compared to the full 31-variable set, demonstrating that the selected 8 variables had a stronger predictive power.Meanwhile, the appearance of AFP_DIFF in the key relevant features also highlights the importance of incorporating dynamic information.
Risk stratification and subgroup analysis
The study identified seven patient subgroups with different clinical characteristics and risk levels based on supervised clustering using SHAP values. Figure B below clearly shows the stratified mortality analysis with gradually increasing mortality probabilities from cluster 1 to cluster 7.
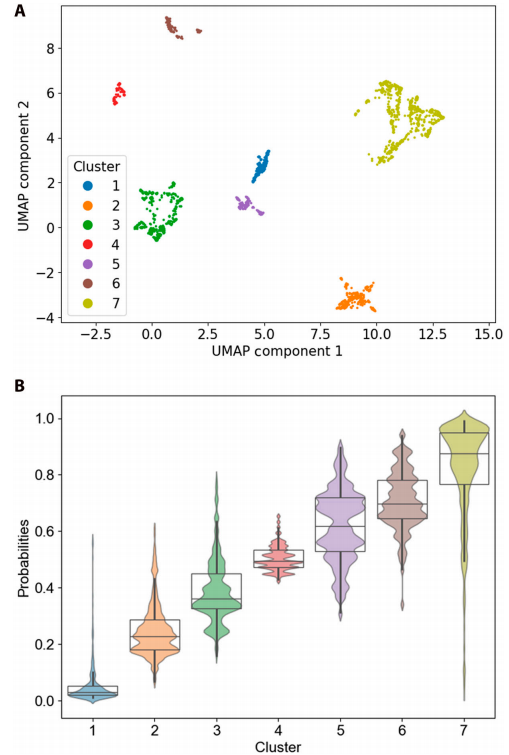
B represents the box plot and population plot of mortality probabilities for 7 clustered observations.
Further analysis based on the Kruskal-Wallis test revealed differences in variables among different clusters. As shown in the SHAP plot, the probability of death increased progressively from cluster 1 to cluster 7; for example, the probability of death for a representative patient increased from 0.03 to 0.98.This trend is consistent with the rankings observed in the box plots, highlighting the effectiveness of the clustering method.
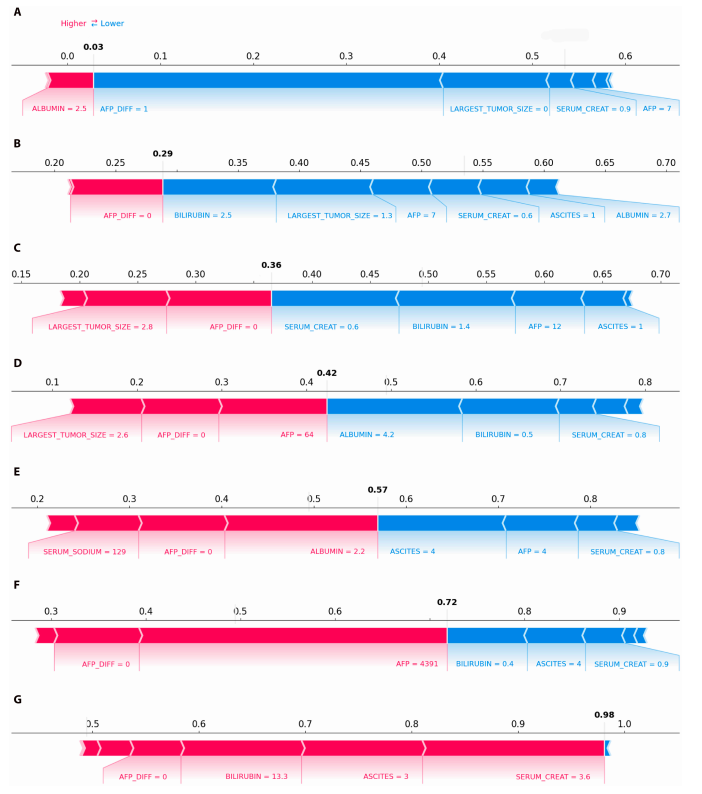
In addition, subgroup analysis clearly revealed two main causes of high mortality risk: severe liver failure (characterized by high bilirubin, high creatinine, and moderate ascites, all of which correspond to positive SHAP values and significantly increase the risk of death) and active tumor progression (characterized by high AFP levels).
In summary, the ELM-HCC machine learning framework based on LightGBM and SHAP interpretability analysis proposed in this study demonstrates significantly better performance than traditional scoring systems in predicting the risk of death during the 3-month waiting period for HCC liver transplant candidates. At the same time, it reveals patient subgroups with different risk characteristics through supervised clustering, providing a more accurate and interpretable risk assessment tool for clinical decision-making.
Innovative methods for risk assessment of liver transplant candidates; a comprehensive approach fills a research gap.
As mentioned above, liver cancer is becoming a global public health challenge. Faced with the increasingly severe disease challenges and higher medical requirements, a scientifically sound and reasonable plan for liver transplant candidate lists is invaluable. As early as 2002, the Model for End-Stage Liver Disease (MELD) was used to prioritize liver transplant candidates. However, after several revisions, the allocation of MELD still cannot fairly satisfy all candidates.
Machine learning, with its ability to process high-dimensional and multimodal data, has now become the best solution for predicting the mortality risk of organ transplant candidates.
Machine learning models have been previously applied to predict liver transplant mortality rates. For example, a joint team from MIT, UC San Francisco, and the University of Texas proposed OPOM, a mortality rate optimization prediction model based on optimal classification trees (OCTs).Based on this model for liver allocation, the number of deaths per year can be reduced by approximately 418 compared to the MELD model, with a significant decrease in the number of deaths/removals across all UNOS regions and disease severity levels.In addition, the model also adjusted the number of livers allocated to patients with and without HCC, which significantly optimized liver transplant allocation and reduced candidate mortality.
Thesis title: Development and validation of an optimized prediction of mortality for candidates awaiting liver transplantation
Paper address:
https://www.sciencedirect.com/science/article/pii/S1600613522090335
However, while OPOM performed well, this model is based on a mixed cohort of HCC and non-HCC patients and does not specifically address the dual risks of liver failure and tumor progression faced by HCC patients. ELM-HCC undoubtedly fills this gap.
Finally, this study not only improves and expands upon previous research, but more importantly, as the authors stated, it fills a gap in current research. By achieving the first-ever interpretable and accurate prediction of mortality during the 3-month waiting period for HCC liver transplant candidates, it provides a new approach to machine learning combined with organ transplant candidate risk assessment.
References:
1. A research team from Telecom Sud-Paris and Paris-Saclay University in France has proposed a machine learning framework that integrates ensemble learning with SHAple Additive exPlanations (SHAP) analysis, providing a new solution for assessing the mortality risk of HCC liver transplant candidates.
2.https://www.sciencedirect.com/science/article/pii/S1600613522090335