Command Palette
Search for a command to run...
dnaHNet:ゲノム配列学習のためのスケーラブルかつ階層的なFoundation Model
dnaHNet:ゲノム配列学習のためのスケーラブルかつ階層的なFoundation Model
Arnav Shah Junzhe Li Parsa Idehpour Adibvafa Fallahpour Brandon Wang Sukjun Hwang Bo Wang Patrick D. Hsu Hani Goodarzi Albert Gu
概要
ゲノム基盤モデルはDNAの構文(syntax)を解読する可能性を秘めているが、その入力表現において根本的なトレードオフに直面している。標準的な固定語彙を用いたtokenizerは、コドンや制御エレメント(regulatory elements)といった生物学的に意味のあるモチーフを断片化させてしまう。一方で、塩基レベル(nucleotide-level)のモデルは生物学的な整合性を維持できるものの、長いコンテキストを扱う際に膨大な計算コストを要するという課題がある。本論文では、ゲノム配列をエンドツーエンドでセグメント化およびモデリングする、最先端のtokenizer-freeな自己回帰モデル「dnaHNet」を提案する。dnaHNetは、微分可能なダイナミックチャンキング(differentiable dynamic chunking)メカニズムを用いることで、生の塩基を潜在的なtokenへと適応的に圧縮し、圧縮率と予測精度のバランスを最適化する。原核生物のゲノムを用いて事前学習を行った結果、dnaHNetはStripedHyena2を含む主要なアーキテクチャを、スケーラビリティと効率性の両面で上回る性能を示した。この再帰的なチャンキングにより、演算量(FLOPs)を二次関数的に削減することが可能となり、Transformerと比較して3倍以上の推論高速化を実現した。ゼロショット(zero-shot)タスクにおいて、dnaHNetはタンパク質変異の適応度(protein variant fitness)や遺伝子の必須性(gene essentiality)の予測で優れた性能を達成したほか、教師なし学習によって階層的な生物学的構造を自動的に発見することに成功した。これらの結果は、dnaHNetが次世代のゲノムモデリングにおける、スケーラブルかつ解釈可能なフレームワークであることを裏付けている。
One-sentence Summary
By employing a differentiable dynamic chunking mechanism to adaptively compress raw nucleotides into latent tokens, the tokenizer-free autoregressive foundation model dnaHNet achieves superior scaling and efficiency over architectures like StripedHyena2 and Transformers, delivering over 3x inference speedup and state-of-the-art performance on zero-shot tasks such as predicting protein variant fitness and gene essentiality.
Key Contributions
- The paper introduces dnaHNet, a tokenizer-free autoregressive model that utilizes a differentiable dynamic chunking mechanism to adaptively compress raw nucleotides into latent tokens.
- This architecture achieves significant computational efficiency by employing recursive chunking to reduce FLOPs quadratically, resulting in over 3x faster inference speeds compared to Transformer models.
- Experiments on prokaryotic genomes demonstrate that the model outperforms leading architectures like StripedHyena2 in scaling and efficiency, while also showing superior zero-shot performance in predicting protein variant fitness and gene essentiality.
Introduction
Genomic foundation models are essential for decoding DNA syntax to advance fields like drug discovery and synthetic biology. However, current approaches face a fundamental tradeoff between computational efficiency and biological accuracy. Fixed-vocabulary tokenizers are efficient but often fragment critical biological motifs like codons, while nucleotide-level models preserve biological coherence but suffer from prohibitive computational costs when processing long genomic contexts. The authors leverage a hierarchical, tokenizer-free architecture called dnaHNet to resolve this tension. By using a differentiable dynamic chunking mechanism, the model adaptively compresses raw nucleotides into latent tokens, allowing it to achieve superior scaling, faster inference, and state-of-the-art performance on zero-shot tasks such as protein variant fitness prediction.
Dataset
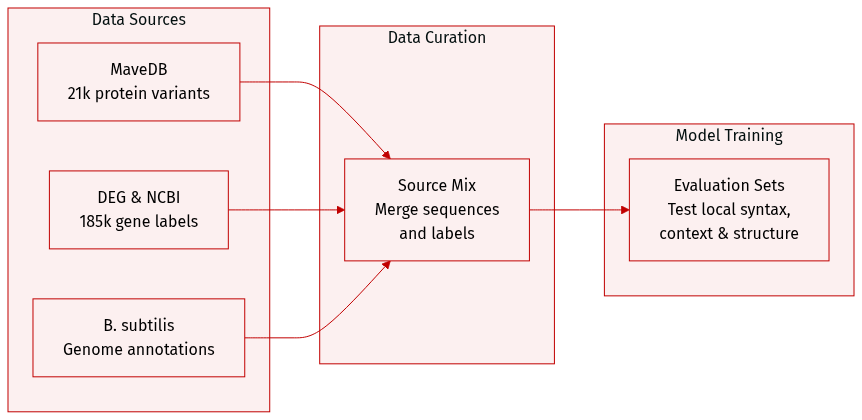
The authors evaluate dnaHNet using three distinct datasets designed to test different biological modeling capabilities:
- Protein Variant Effects (MaveDB): This subset consists of 21,250 nucleotide-level data points compiled from 12 experimental fitness datasets for E. coli K-12. It is used to assess the model's ability to capture local coding syntax and predict protein fitness landscapes.
- Gene Essentiality (DEG): Comprising 185,226 data points, this dataset was constructed by generating binary essentiality labels for genes across 62 bacterial organisms from the Database of Essential Genes. The authors sourced base sequences and annotations from NCBI and labeled genes as essential if they matched DEG entries by name or sequence identity greater than 99 percent. This subset evaluates the model's capacity to integrate genomic context and long-range dependencies.
- Genomic Structure Interpretation (NCBI): To perform interpretability analysis, the authors used the B. subtilis genome and functional annotations from NCBI. They partitioned the genome into distinct functional regions based on these annotations to determine how the model's segmentation aligns with known biological structures.
Method
The authors introduce dnaHNet, a scalable, tokenizer-free foundation model designed for genomic sequence learning. The model formulates genomic learning as an autoregressive sequence modeling problem. Given a nucleotide sequence X=(x1,…,xL) where xt∈{A,C,G,T}, the objective is to model the probability distribution P(X)=∏t=1LP(xt∣x<t). To handle long genomic contexts efficiently, dnaHNet utilizes a recursive hierarchical architecture consisting of three primary differentiable modules: an Encoder (E), a Main Network (M), and a Decoder (D).
Refer to the framework diagram:
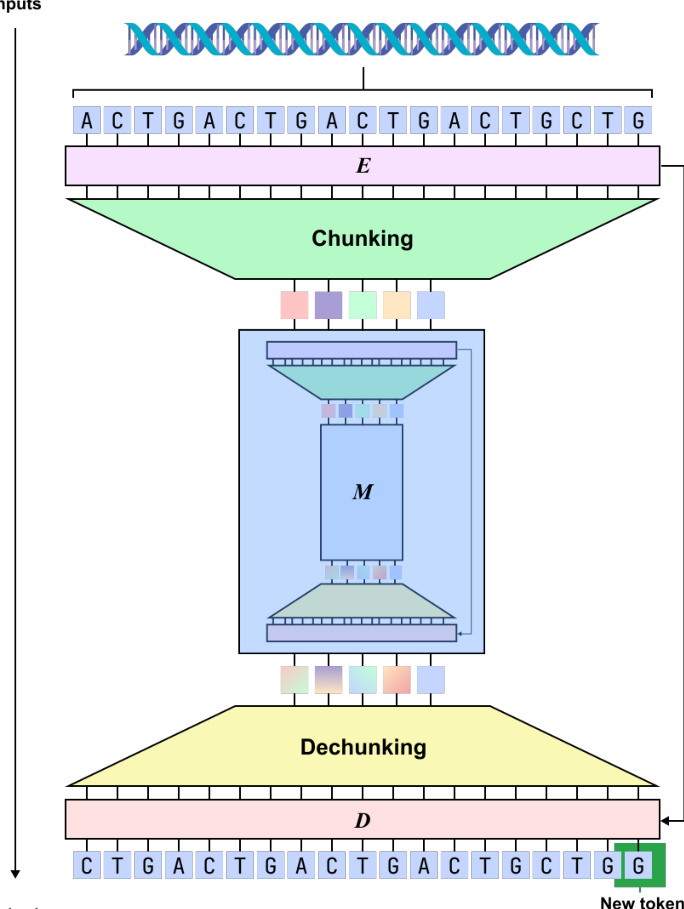
The Encoder is responsible for compressing nucleotide-level inputs into latent chunks through a dynamic segmentation mechanism. It employs a hybrid backbone consisting of four Mamba layers and one Transformer layer. The Encoder transforms input embeddings into hidden states h1:L∈RL×D. To determine segmentation boundaries, a boundary prediction module computes probabilities pt∈[0,1] using the following formulation: \npt=21(1−CosineSim(Wqht,Wkht−1)) where Wq and Wk are learnable projection matrices. High boundary probabilities are assigned to nucleotides with dissimilar representations, which encourages the model to segment at biologically significant transitions, such as codon boundaries. The Chunking layer then downsamples the output by selecting representations at these predicted boundaries, resulting in a compressed sequence E=(e1,…,eL′) where L′≤L.
As shown in the figure below:
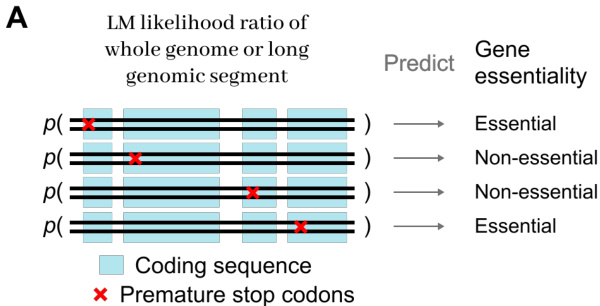
The compressed sequence E is then processed by the Main Network M. This module can be a standard Transformer or another H-Net module, which allows for recursive chunking to capture multiple levels of abstraction. In a two-stage hierarchy, the first stage captures high-frequency local patterns like codon periodicity, while the second stage models long-range dependencies across functional regions. The Main Network outputs processed latent states E^=(e^1,…,e^L′)∈RL′×D.
The Decoder D maps these latent states back to the original nucleotide resolution through a two-step process. First, a smoothing module refines the latent states into smoothed representations Eˉ=(eˉ1,…,eˉL′) using a recurrence that interpolates discrete chunks: eˉj=Pje^j+(1−Pj)eˉj−1 where Pj is the boundary probability for the j-th chunk. Second, an upsampler expands these smoothed latents to the original length L by copying the vector eˉc(t) to every nucleotide position t corresponding to the chunk index c(t). The Decoder then utilizes four Mamba layers and one Transformer layer to model autoregressive dependencies, with a linear head projecting the output to the nucleotide vocabulary logits to produce the next-nucleotide distribution.
The training process is conducted end-to-end using a composite objective. The primary component is the autoregressive next-token prediction loss: LNLL=−∑t=1LlogPθ(xt∣x<t) To prevent degenerate segmentation during training, the authors incorporate a ratio loss that regularizes the dynamic chunking toward a target downsampling ratio Rs for each stage s: Lrate(s)=Rs−1Rs((Rs−1)FsGs+(1−Fs)(1−Gs)) where Fs is the actual fraction of selected chunks and Gs is the average boundary probability. The total loss is defined as L=LNLL+α∑sLrate(s), where α is a regularization coefficient.
Experiment
The experiments compare dnaHNet against StripedHyena2 and Transformer++ architectures through scaling law analyses, zero-shot biological prediction tasks, and structural interpretability studies. Results demonstrate that dnaHNet achieves superior compute and inference efficiency, consistently outperforming baselines in perplexity scaling and downstream performance on protein variant effect and gene essentiality predictions. Furthermore, the model demonstrates an emergent ability to learn biological hierarchies without supervision, effectively discovering codon structures and functional genomic regions through its hierarchical compression mechanism.
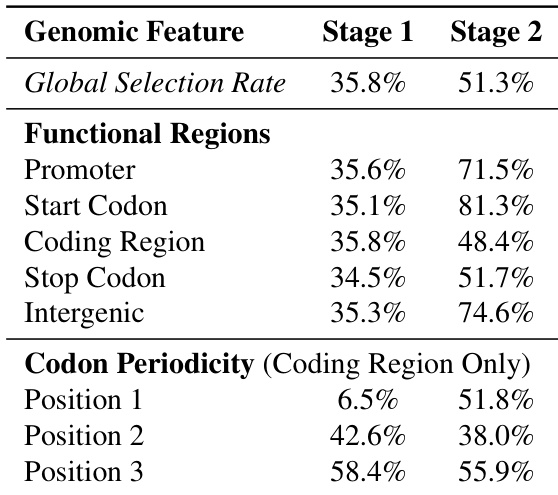
The authors analyze the learned hierarchical chunking boundaries of the dnaHNet model across different stages. The results demonstrate that the first stage captures local triplet codon structure, while the second stage identifies broader functional genomic organization. The first stage shows strong periodicity within coding regions, with selection rates varying significantly by codon position. The second stage exhibits higher selection rates for functional regions like promoters and intergenic areas compared to coding regions. The overall global selection rate increases from the first stage to the second stage.
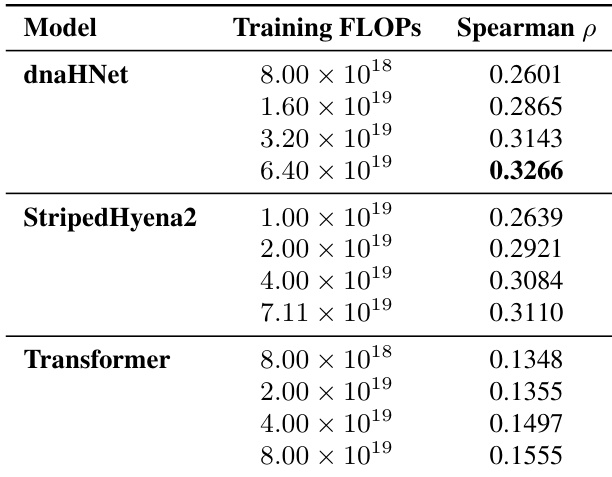
The authors evaluate the zero-shot protein variant effect prediction performance of dnaHNet against StripedHyena2 and Transformer baselines across various training compute budgets. Results show that dnaHNet consistently achieves higher Spearman correlation for predicting experimental fitness as training FLOPs increase. dnaHNet demonstrates superior predictive accuracy compared to both StripedHyena2 and Transformer architectures. The predictive performance of dnaHNet improves steadily as the amount of training compute increases. The performance gap between dnaHNet and the baseline models widens at higher training compute scales.
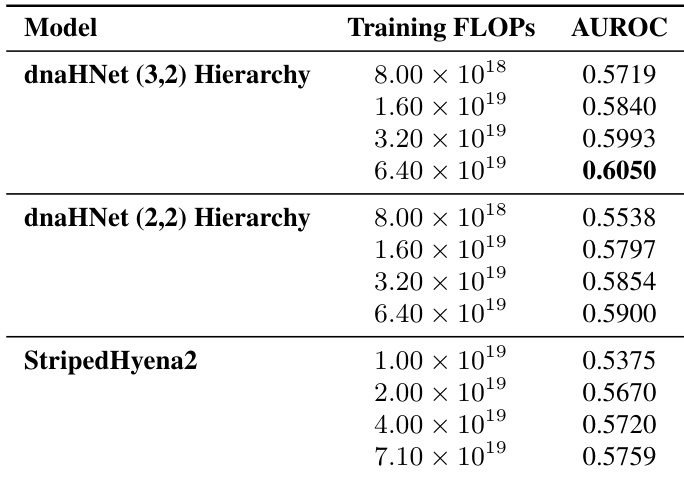
The authors compare the gene essentiality prediction performance of two dnaHNet hierarchical configurations against a StripedHyena2 baseline across various compute budgets. Results show that both dnaHNet architectures consistently outperform the baseline in AUROC as training compute increases. Both dnaHNet hierarchy versions demonstrate improved predictive accuracy as training FLOPs increase. The dnaHNet (3,2) hierarchy achieves higher AUROC scores compared to the (2,2) hierarchy at matched compute levels. dnaHNet models maintain a performance advantage over StripedHyena2 across the tested compute range.
The researchers evaluate the dnaHNet model by analyzing its hierarchical chunking boundaries and testing its predictive capabilities for protein variant effects and gene essentiality. The findings reveal that the model successfully captures both local codon structures and broader functional genomic organization through its multi-stage hierarchy. Across various compute budgets, dnaHNet consistently outperforms StripedHyena2 and Transformer baselines, demonstrating superior scaling and predictive accuracy in both protein fitness and gene essentiality tasks.