Command Palette
Search for a command to run...
ScribblePrompt: あらゆる生体医用画像に対する高速かつ柔軟なインタラクティブ・セグメンテーション
ScribblePrompt: あらゆる生体医用画像に対する高速かつ柔軟なインタラクティブ・セグメンテーション
Halle E. Wong Marianne Rakic John Guttag Adrian V. Dalca
概要
バイオメディカル画像セグメンテーションは、科学研究と臨床ケアの両面において極めて重要な要素である。十分なラベル付きデータがあれば、ディープラーニングモデルを学習させることで、特定のバイオメディカル画像セグメンテーションタスクを正確に自動化することが可能となる。しかし、学習データを作成するために手動で画像をセグメンテーションする作業は、非常に労力を要し、かつ専門知識を必要とする。本稿では、バイオメディカルイメージング向けに、ニューラルネットワークに基づいた柔軟なインタラクティブ・セグメンテーション・ツールである「ScribblePrompt」を提案する。ScribblePromptを使用することで、アノテーターはscribbles(スクリブル)、clicks(クリック)、およびbounding boxes(バウンディングボックス)を用いて、未知の構造に対してもセグメンテーションを行うことが可能となる。厳密な定量実験を通じて、同等のインタラクション回数において、ScribblePromptは学習時に含まれていないデータセットに対し、従来手法よりも正確なセグメンテーションを実現することを実証した。また、専門家を対象としたユーザー調査では、ScribblePromptは既存の最良の手法と比較して、Dice係数を15%向上させつつ、アノテーション時間を28%削減することに成功した。ScribblePromptの成功は、一連の綿密な設計上の決定に基づいている。これには、極めて多様な画像セットとタスクを組み込んだ学習戦略、シミュレートされたユーザーインタラクションおよびラベルのための新規アルゴリズム、そして高速な推論を可能にするネットワークが含まれる。我々は、インタラクティブなデモを通じてScribblePromptの動作を公開するとともに、コードおよびscribbleアノテーションのデータセットを https://scribbleprompt.csail.mit.edu で公開している。
One-sentence Summary
Researchers from MIT propose ScribblePrompt, a flexible neural network-based interactive segmentation tool that utilizes scribbles, clicks, and bounding boxes to segment unseen biomedical structures, outperforming previous methods by reducing expert annotation time by 28% and improving Dice scores by 15% through a diverse training strategy and novel simulated interaction algorithms.
Key Contributions
- The paper introduces ScribblePrompt, a flexible neural network-based interactive segmentation framework that supports multiple user inputs including scribbles, clicks, and bounding boxes. This method allows for the segmentation of previously unseen biomedical structures at inference time without the need for task-specific retraining.
- The work presents novel algorithms for simulating realistic user interactions and generating synthetic labels, which facilitates training on a highly diverse set of images and tasks. This simulation engine enables the model to generalize effectively to new datasets and specialized medical imaging modalities.
- Experimental results and user studies demonstrate that the system outperforms previous methods by achieving a 15% improvement in Dice score and a 28% reduction in annotation time compared to the next best baseline. The ScribblePrompt-UNet architecture also provides computational efficiency capable of running on a CPU.
Introduction
Accurate biomedical image segmentation is essential for clinical care and scientific research, yet manual annotation remains a labor intensive process requiring significant domain expertise. Existing deep learning methods often struggle with generalization, as they are typically trained for specific tasks or modalities and fail when encountering unseen structures. While vision foundation models like SAM show promise, they often perform poorly on subtle biomedical delineations and require limited interaction types. The authors leverage a new framework called ScribblePrompt to enable flexible, interactive segmentation across diverse biomedical images using scribbles, clicks, and bounding boxes. By introducing a novel scribble simulation engine and a diverse training strategy, the authors provide a tool that generalizes to unseen tasks without retraining, significantly reducing annotation time while improving segmentation accuracy.
Dataset
The authors developed a comprehensive biomedical imaging framework using the following data strategies:
- Dataset Composition and Sources: The training collection is built upon large scale efforts like MegaMedical, comprising 77 open access biomedical imaging datasets. This collection includes over 54,000 scans across 16 image types and 711 labels, spanning diverse domains such as the brain, thorax, abdomen, spine, cells, skin, eyes, and more.
- Task Definition and Processing:
- 2D segmentation tasks are defined by a combination of the dataset, the specific axis (for 3D modalities), and the label.
- For datasets with multiple labels, each is treated as a separate binary segmentation task.
- For 3D volumes, the authors extract the middle slice and the slice containing the maximum label area.
- To prevent overfitting, the authors implement a synthetic label mechanism where a superpixel algorithm partitions an image into a multi-label mask, from which a single label is randomly selected to replace the ground truth with a specific probability.
- Training Strategy: The authors use hierarchical sampling during training to balance datasets of different sizes. This sampling is performed by dataset and modality, then by axis, and finally by label. Both the input images and the sampled segmentations undergo data augmentation before simulating user interactions.
- MedScribble Dataset: For manual evaluation, the authors curated the MedScribble dataset, which contains manual scribble annotations from three annotators for 64 image segmentation pairs. These pairs were randomly selected from the validation splits of 14 different datasets. A specific subset of 31 image segmentation pairs from 7 unseen datasets is used to report results for manual scribble evaluation.
Method
The ScribblePrompt framework is designed as an interactive segmentation method that generalizes across diverse biomedical imaging modalities. The core objective is to learn a function fθ(xt,ui,y^i−1t) that produces an iterative segmentation y^i given an input image xt, a set of user interactions ui, and the previous prediction y^i−1t. The model is optimized by minimizing the difference between the true segmentation yt and the k iterative predictions through a supervised segmentation loss:
L(θ;T)=Et∈T[E(xt,yt)∈t[∑i=1kLSeg(yt,ftheta(xt,ui,y^i−1t))]]
The training process involves simulating a sequence of interactive steps. As shown in the framework diagram:
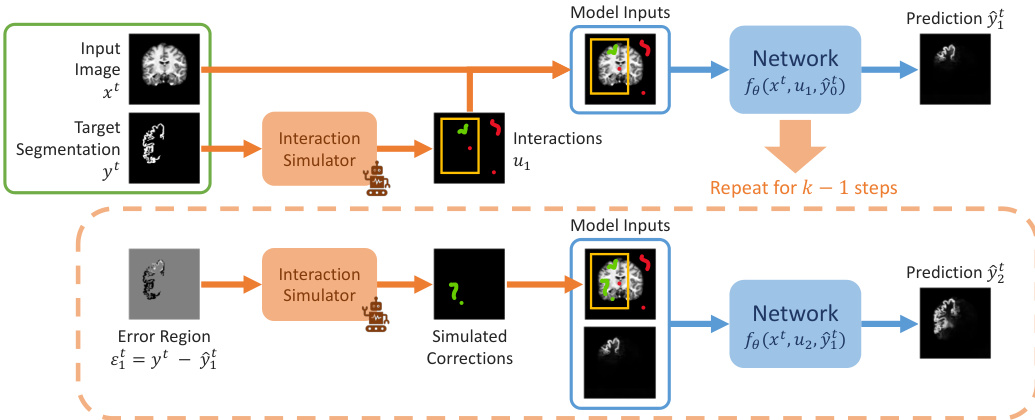
Initially, a set of interactions u1, such as bounding boxes, clicks, or scribbles, is simulated based on the ground truth yt to produce the first prediction y^1t. In subsequent steps, the framework simulates user corrections by identifying the error region εit=yt−y^it and generating new interactions ui+1 based on this error. This iterative process repeats for k steps to refine the segmentation.
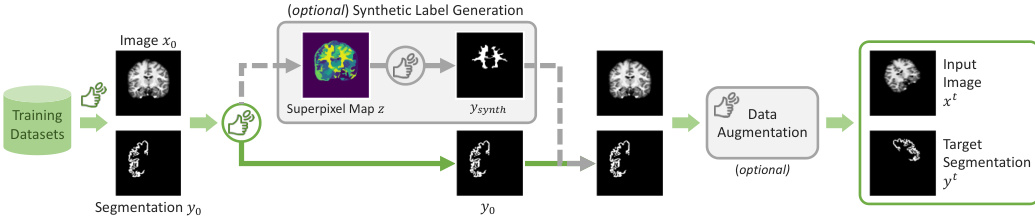
To enhance generalization and prevent the model from overfitting to specific tasks, the authors incorporate a synthetic label generation mechanism. During training, a sample (x0,y0) may be replaced by a synthetic label ysynth with a probability psynth. This is achieved by applying a superpixel algorithm to the image x0 to create a map of k superpixels, from which a single superpixel is randomly selected to serve as the synthetic target. The overall training flow, including this optional augmentation, is illustrated in the figure below:
The interaction simulation utilizes several strategies for different prompt types. For scribbles, the authors implement line, centerline, and contour strategies, which are then corrupted through random masking and deformation to mimic human variability. For clicks, they employ random, center, or interior border region sampling. Bounding boxes are simulated by computing the minimum enclosing box of the label and enlarging it slightly. These prompts are encoded into input channels, allowing the network to process them efficiently. For the ScribblePrompt-UNet architecture, the input consists of five channels: the image, bounding box encoding, positive click/scribble encoding, negative click/scribble encoding, and the previous prediction logits.
Experiment
The researchers evaluated ScribblePrompt through manual scribble tests, simulated iterative interactions, a user study with experienced neuroimaging researchers, and computational runtime analysis. The experiments validate the model's ability to generalize to unseen medical modalities and anatomical regions using flexible prompts such as bounding boxes, clicks, and scribbles. Findings demonstrate that ScribblePrompt provides superior segmentation accuracy and greater efficiency than existing generalist models, offering a more responsive and user-friendly experience that significantly reduces annotation time.
The authors compare ScribblePrompt-UNet against the SAM (ViT-b) model through a user study involving experienced annotators. Results indicate that ScribblePrompt-UNet achieves higher segmentation accuracy and lower error rates while requiring less time and fewer interactions per task. ScribblePrompt-UNet achieves a higher mean Dice score compared to SAM (ViT-b). The ScribblePrompt-UNet model results in lower HD95 values than the SAM baseline. Users completed tasks more efficiently with ScribblePrompt-UNet, requiring less time and fewer interaction steps per task.

The authors evaluate the ScribblePrompt models against several baseline methods using manual scribble inputs across different datasets. Results show that both ScribblePrompt-UNet and ScribblePrompt-SAM achieve higher Dice scores and lower Hausdorff Distance compared to existing interactive segmentation methods. ScribblePrompt models outperform SAM variants and MedSAM when using manual scribble prompts The ScribblePrompt models demonstrate superior accuracy and boundary adherence compared to MIDeepSeg ScribblePrompt variants achieve the highest Dice scores among all compared methods in the manual scribble evaluation
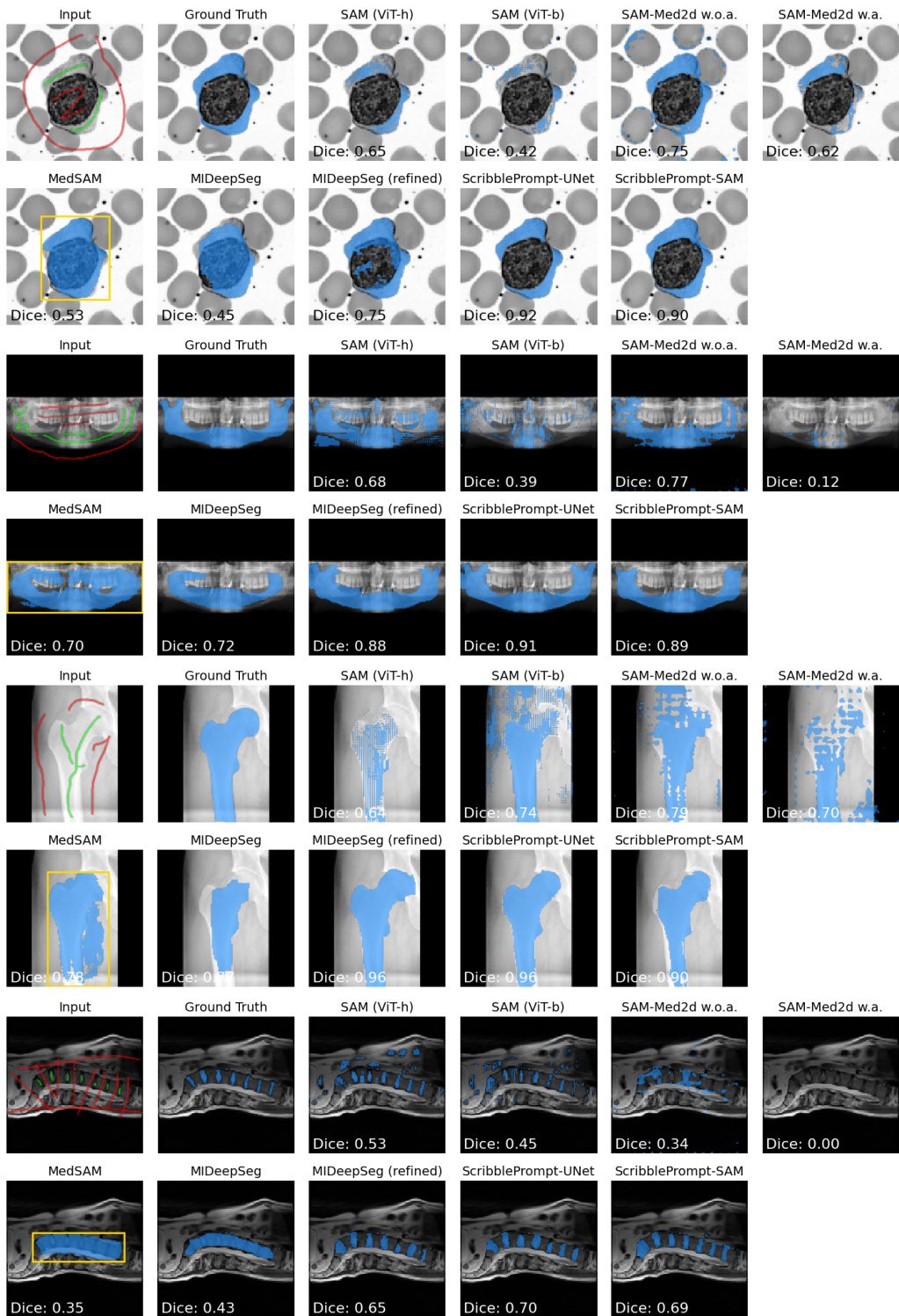
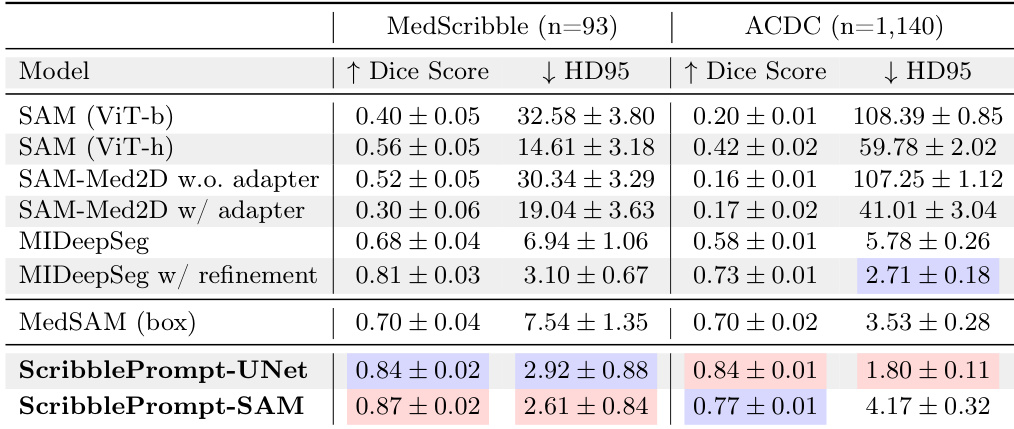
The authors evaluate several interactive segmentation models using manual scribbles on the MedScribble and ACDC datasets. Results show that the ScribblePrompt models achieve the highest Dice scores and the lowest Hausdorff Distance compared to all other tested methods. ScribblePrompt-SAM and ScribblePrompt-UNet outperform existing methods like SAM, SAM-Med2D, and MedSAM in segmentation accuracy. ScribblePrompt-UNet achieves the best overall performance on the ACDC dataset in terms of both Dice score and Hausdorff Distance. The ScribblePrompt models demonstrate superior ability to handle manual scribble inputs compared to SAM-based baselines.
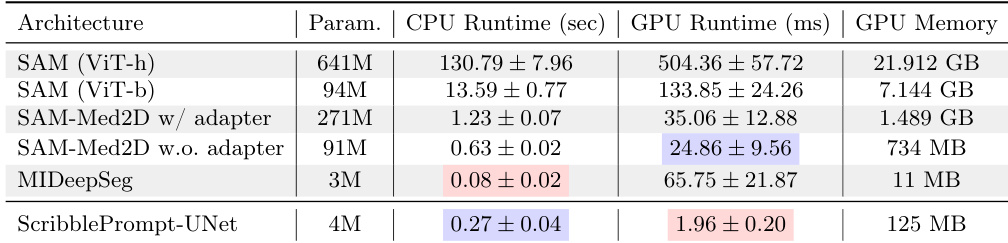
The authors compare the computational efficiency of ScribblePrompt-UNet against several SAM-based models and MIDeepSeg. Results show that ScribblePrompt-UNet maintains low latency on both CPU and GPU hardware. ScribblePrompt-UNet achieves significantly faster GPU inference times compared to all other evaluated models. On a single CPU, ScribblePrompt-UNet demonstrates competitive performance with low latency, outperforming larger SAM variants. The models show a clear trade-off between parameter count and inference speed, with ScribblePrompt-UNet providing high efficiency with a small parameter footprint.
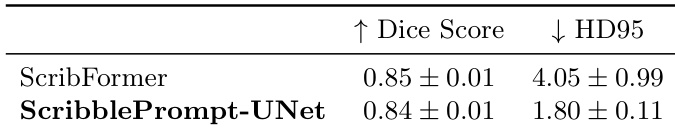
The authors compare ScribblePrompt-UNet to ScribFormer, a scribble-supervised learning method, using the ACDC dataset. Results show that ScribblePrompt-UNet achieves a comparable Dice score while maintaining a significantly lower HD95. ScribblePrompt-UNet achieves a Dice score similar to ScribFormer ScribblePrompt-UNet demonstrates much better boundary accuracy as indicated by a lower HD95 The performance of ScribblePrompt-UNet is competitive with specialized scribble-supervised models
The authors evaluated ScribblePrompt models through user studies, comparative segmentation benchmarks on MedScribble and ACDC datasets, and computational efficiency tests. The results demonstrate that ScribblePrompt models provide superior segmentation accuracy and better boundary adherence than existing SAM variants and specialized scribble-supervised methods. Furthermore, the models enhance user productivity by requiring fewer interactions and offer high computational efficiency with low latency across different hardware.