Command Palette
Search for a command to run...
Generative AI Enables Structural Brain Network Construction from fMRI via Symmetric Diffusion Learning
Generative AI Enables Structural Brain Network Construction from fMRI via Symmetric Diffusion Learning
Qiankun Zuo Bangjun Lei Wanyu Qiu Changhong Jing Jin Hong Shuqiang Wang
Abstract
Mapping from functional connectivity (FC) to structural connectivity (SC) can facilitate multimodal brain network fusion and discover potential biomarkers for clinical implications. However, it is challenging to directly bridge the reliable non-linear mapping relations between SC and functional magnetic resonance imaging (fMRI). In this paper, a novel symmetric diffusive generative adversarial network-based fMRI-to-SC (DiffGAN-F2S) model is proposed to predict SC from brain fMRI in a unified framework. To be specific, the proposed DiffGAN-F2S leverages denoising diffusion probabilistic models (DDPMs) and adversarial learning to efficiently generate symmetric and high-fidelity SC through a few steps from fMRI. By designing the dual-channel multi-head spatial attention (DMSA) and graph convolutional modules, the symmetric graph generator first captures global relations among direct and indirect connected brain regions, then models the local brain region interactions. It can uncover the complex mapping relations between fMRI and symmetric structural connectivity. Furthermore, the spatially connected consistency loss is devised to constrain the generator to preserve global-local topological information for accurate symmetric SC prediction. Testing on the public Alzheimer's Disease Neuroimaging Initiative (ADNI) dataset, the proposed model can effectively generate empirical SC-preserved connectivity from four-dimensional imaging data and shows superior performance in SC prediction compared with other related models. Furthermore, the proposed model can identify the vast majority of important brain regions and connections derived from the empirical method, providing an alternative way to fuse multimodal brain networks and analyze clinical brain disease.
One-sentence Summary
The authors propose DiffGAN-F2S, a symmetric diffusion generative adversarial network that directly predicts structural connectivity from fMRI in one stage. By integrating dual-channel attention and a spatially connected consistency loss, this model outperforms prior methods in accuracy and efficiency for analyzing neurodegenerative diseases like Alzheimer's.
Key Contributions
- The paper introduces DiffGAN-F2S, a symmetric diffusive generative adversarial network that predicts structural connectivity from fMRI data by integrating denoising diffusion probabilistic models with adversarial learning to generate high-fidelity graphs in a unified framework.
- A dual-channel multi-head spatial attention mechanism combined with graph convolutional modules is designed to capture global relations among direct and indirect brain regions while modeling local interactions to uncover complex non-linear mapping relations.
- Experiments on the ADNI dataset demonstrate that the method effectively preserves empirical connectivity patterns and outperforms existing models in structural connectivity prediction, successfully identifying critical brain regions and connections for clinical analysis.
Introduction
Modeling the relationship between functional connectivity derived from fMRI and structural connectivity is vital for uncovering pathological mechanisms in neurodegenerative diseases like Alzheimer's. Prior approaches often rely on inefficient two-stage pipelines that first compute functional connectivity before predicting structure, while also suffering from training instability and mode collapse in generative models. The authors leverage a novel symmetric diffusive generative adversarial network called DiffGAN-F2S to predict structural connectivity directly from fMRI in a unified, end-to-end framework. By integrating denoising diffusion probabilistic models with adversarial learning and a dual-channel multi-head spatial attention mechanism, their method efficiently generates high-fidelity symmetric graphs while preserving critical global and local topological information.
Dataset
-
Dataset Composition and Sources: The authors utilize data from the Alzheimer's Disease Neuroimaging Initiative (ADNI) public repository, selecting 240 patients divided equally into two categories: 120 normal controls and 120 subjects with mild cognitive impairment (MCI), which includes both early and late stages.
-
Key Details for Each Subset:
- Imaging Modalities: Every subject underwent scanning using a 3T magnetic resonance instrument for both functional MRI (fMRI) and Diffusion Tensor Imaging (DTI).
- DTI Specifications: Acquisition parameters included a repetition time (TR) of 3.4 s to 17.5 s, an echo time (TE) of 56 ms to 105 ms, and 6 to 126 gradient directions.
- fMRI Specifications: The TR varied between 0.607 s and 3.0 s, while the TE ranged from 30 ms to 32 ms.
-
Data Processing and Preprocessing:
- DTI Pipeline: The authors processed DTI data using the PANDA toolbox and the anatomical automatic labeling (AAL90) atlas to generate an empirical structural connectivity (SC) matrix (A0) with dimensions of 90×90.
- fMRI Pipeline: Instead of standard procedures, the team used the
aal.niiatlas file to convert raw fMRI data into ROI-based time series without additional parameters, resulting in a primary sample matrix (F) of size 90×187. - Matrix Definition: In the 90×187 fMRI matrix, the 90 rows correspond to the ROIs defined by the AAL90 atlas, and each element represents the mean signal intensity of a specific ROI at a given time point.
-
Model Usage: The processed datasets serve as the foundation for testing the model's performance in predicting structural connectivities and comparing them against empirical SCs across different diffusive steps and subject groups.
Method
The authors propose the DiffGAN-F2S model to predict Structural Connectivity (SC) from brain fMRI data. The overall framework is illustrated in the first image, which depicts the transformation from raw brain data to fMRI, processed by the proposed model to yield structural connectivity.
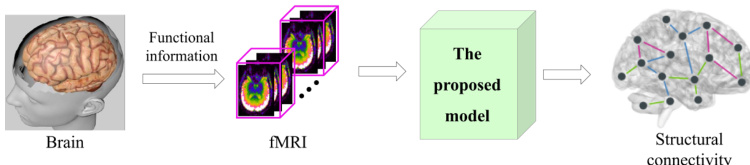
The architecture consists of two primary components: a symmetric diffusive process and a conditional denoising process. As shown in the figure below, the diffusive process models the truth distribution of noisy SC by persistently injecting Gaussian noise into the empirical SC matrix A0 over T steps until it becomes a Gaussian matrix AT.
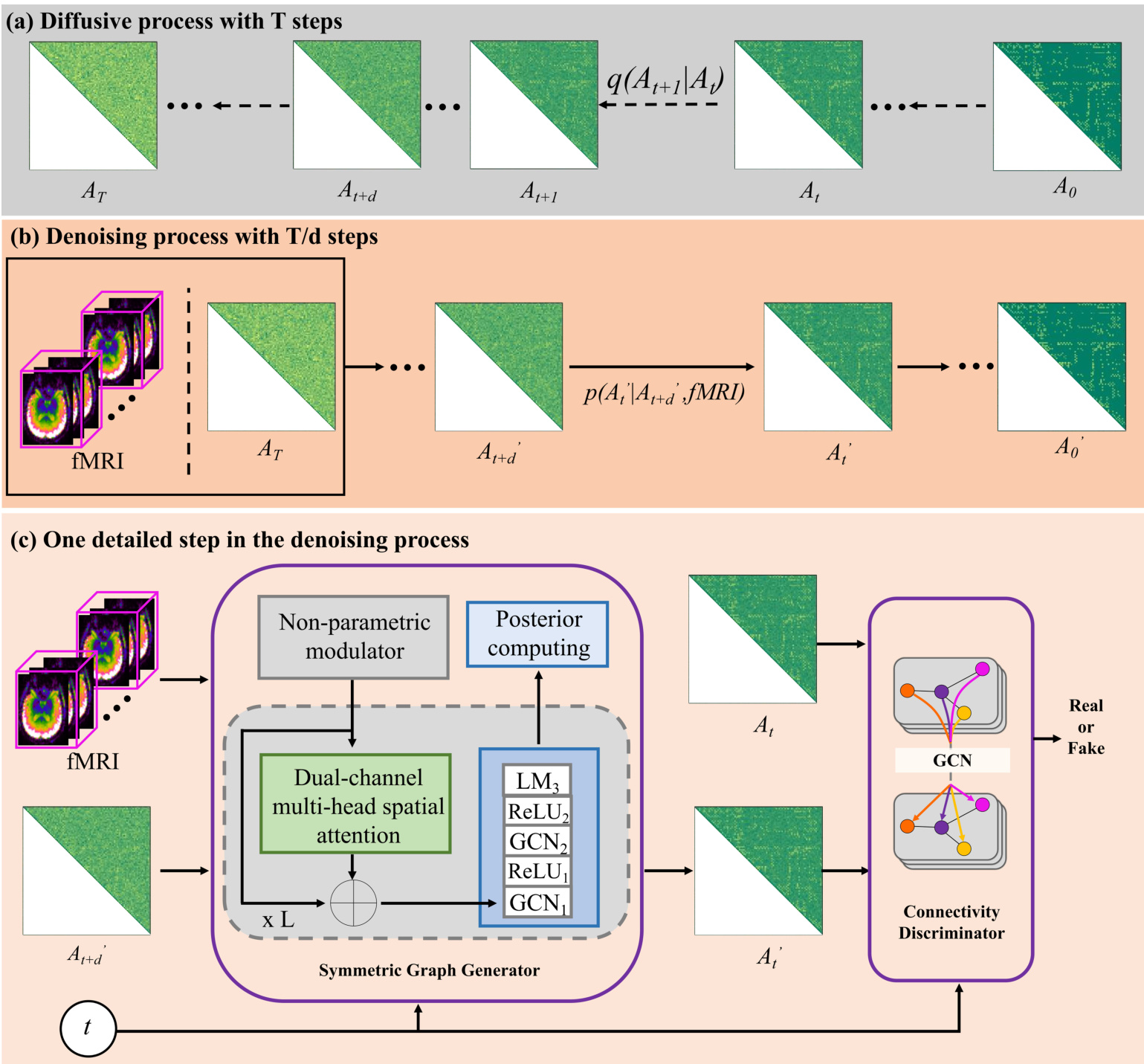
In the denoising process, the brain fMRI serves as a condition to guide the prediction of the clean SC A0′. To reduce computational cost, the denoising steps are compressed to T/d steps, where each step utilizes a conditional generative adversarial network. The detailed structure of a single denoising step involves a Symmetric Graph Generator and a Connectivity Discriminator.
The Symmetric Graph Generator employs a Non-parametric modulator (NPM) to transform raw fMRI into preliminary ROI-based time series without learnable parameters. These features, along with the noisy sample, pass through L layers containing a Dual-channel multi-head spatial attention (DMSA) module and GCN-based modules. The DMSA module is designed to capture global relations among directly and indirectly connected brain regions. The specific implementation of this attention mechanism is detailed in the following figure.
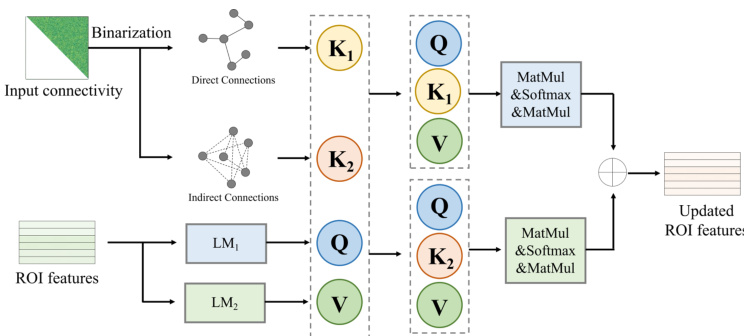
The module computes Query (Q), Key (K1,K2), and Value (V) vectors, where K1 and K2 represent direct and indirect connected ROI features derived from one-layer GCNs. The attention values are calculated separately for direct and indirect connections and combined to update the ROI features. Following the attention mechanism, a Posterior Computing Module (PCD) translates the features into a symmetrically blurred A˙0 and predicts the noisy SC at the current step via posterior sampling.
The Connectivity Discriminator distinguishes whether the noisy SC originates from the generator or the empirical method. It utilizes three GCN layers where the edge represents the noisy SC and the node feature is a one-hot vector corresponding to the 90 brain regions defined by the AAL90 atlas. Temporal embeddings are inserted as bias terms into the discriminating computation.
To optimize the model, three loss functions are devised: the denoising adversarial loss to constrain distribution consistency, the mean absolute error (MAE) loss to measure edge strength differences, and the spatially connected consistency loss to capture local and global graphical properties. The spatially connected consistency loss includes terms for overall similarity and betweenness centrality to ensure accurate prediction of intrinsic structural patterns.
Experiment
- The model transforms fMRI into structural connectivity in a single stage, validating its ability to denoise Gaussian noise into empirical connectivity while preserving both global and local patterns.
- Comparative experiments against seven existing models demonstrate superior accuracy and topological consistency, with predicted results showing high correlation to empirical data and significantly lower errors in graph metrics.
- Connectivity analysis confirms the model reliably identifies disease-relevant brain regions and abnormal connections associated with Mild Cognitive Impairment, matching empirical findings in the majority of key regions.
- Ablation studies verify that the adversarial loss, spatially connected consistency loss, symmetric design, and specific attention mechanisms are critical for maintaining distribution consistency and capturing global topological features.
- Efficiency tests show that the skipping step mechanism balances prediction speed and accuracy, reducing inference time to approximately 3 seconds without significant loss in performance.