Command Palette
Search for a command to run...
BioInstruct: Instruction Tuning von Large Language Models für die biomedizinische natürliche Sprachverarbeitung
BioInstruct: Instruction Tuning von Large Language Models für die biomedizinische natürliche Sprachverarbeitung
Hieu Tran Zhichao Yang Zonghai Yao Hong Yu
Zusammenfassung
Hier ist die professionelle Übersetzung des Textes ins Deutsche, unter Berücksichtigung der wissenschaftlichen Standards und Ihrer spezifischen Vorgaben:„Ziel dieser Arbeit ist es, die Leistungsfähigkeit von Large Language Models (LLMs) im Bereich der biomedizinischen natürlichen Sprachverarbeitung (BioNLP) zu steigern, indem ein domänenspezifischer Instruction-Datensatz eingeführt und dessen Auswirkungen in Kombination mit Prinzipien des Multi-Task Learning untersucht werden. Wir haben BioInstruct erstellt, einen Datensatz bestehend aus 25.005 Instructions für das Instruction-Tuning von LLMs (LLaMA 1 & 2, Versionen 7B & 13B). Die Instructions wurden generiert, indem das Sprachmodell GPT-4 mittels drei zufällig aus 80 von Menschen kuratierten Instructions gezogenen Seed-Samples (Beispielen) angewiesen wurde. Für das parametereffiziente Fine-Tuning haben wir Low-Rank Adaptation (LoRA) eingesetzt. Anschließend evaluierten wir diese instruction-tuned LLMs anhand mehrerer BioNLP-Aufgaben, die in drei Hauptkategorien unterteilt werden können: Question Answering (QA), Information Extraction (IE) und Text Generation (SEN). Zudem untersuchten wir, ob die Kategorien der Instructions (z. B. QA, IE und Generation) die Modellleistung beeinflussen. Im Vergleich zu LLMs ohne Instruction-Tuning zeigten unsere instruction-tuned LLMs signifikante Leistungssteigerungen: durchschnittlich 17,3 % bei der QA-Metrik (Accuracy), 5,7 % bei der IE-Metrik (F1-Score) und 96 % bei den Generation-Aufgaben (GPT-4 Score). Unser instruction-tuned LLaMA 1 Modell mit 7B Parametern war mit anderen LLMs im biomedizinischen Bereich wettbewerbsfähig oder übertraf diese sogar, selbst wenn diese ebenfalls auf Basis von LLaMA 1 mit umfangreichen domänenspezifischen Daten oder einer Vielzahl von Aufgaben feinabgestimmt worden waren.“
One-sentence Summary
To enhance large language models for biomedical natural language processing, the authors propose BioInstruct, a domain-specific dataset of 25,005 instructions generated via GPT-4 that, when used to instruction-tune LLaMA 1 and 2 (7B and 13B) models using Low-Rank Adaptation (LoRA), significantly improves performance across question answering, information extraction, and text generation tasks.
Key Contributions
- The paper introduces BioInstruct, a domain-specific instruction dataset consisting of 25,005 instructions generated by prompting GPT-4 with samples drawn from human-curated biomedical tasks.
- This work implements parameter-efficient fine-tuning on LLaMA 1 and LLaMA 2 models using Low-Rank Adaptation (LoRA) to adapt large language models to the biomedical natural language processing domain.
- Experimental results demonstrate that the instruction-tuned models achieve significant performance gains across question answering, information extraction, and text generation tasks, with the 7B-parameter LLaMA 1 model performing competitively against other domain-specific models.
Introduction
Large language models (LLMs) are increasingly vital for biomedical natural language processing (BioNLP) tasks such as medical question answering and information extraction. While traditional fine-tuning on task-specific datasets can achieve high accuracy, it often requires massive amounts of high-quality annotated data and is prone to overfitting. The authors address these challenges by introducing BioInstruct, a domain-specific instruction tuning dataset consisting of 25,005 instructions generated via GPT-4. By applying parameter-efficient fine-tuning with Low-Rank Adaptation (LoRA) on LLaMA models, the authors demonstrate significant performance gains across question answering, information extraction, and text generation tasks.
Dataset
-
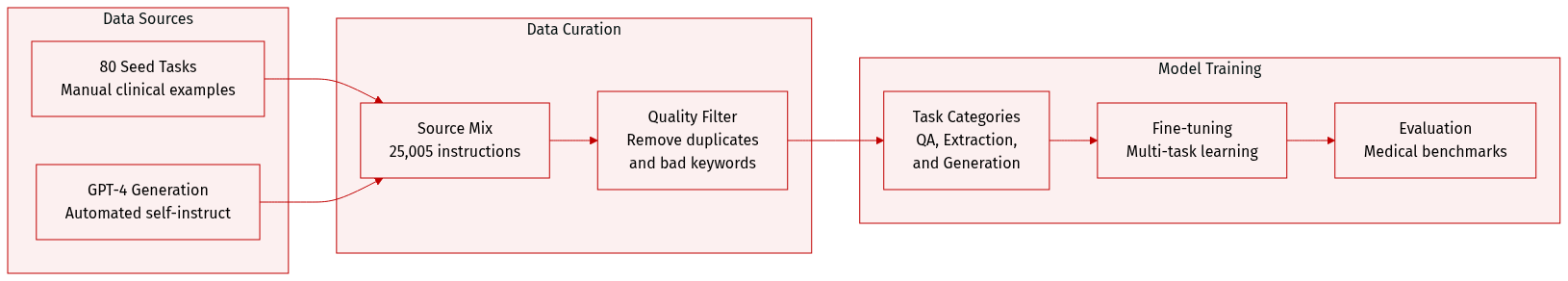
Dataset Composition and Sources: The authors introduce BioInstruct, a dataset consisting of 25,005 natural language instructions designed for biomedical and clinical NLP tasks. The dataset was generated using an automated "Self-Instruct" methodology, starting from 80 manually constructed seed tasks that cover diverse areas such as biomedical question answering, summarization, clinical trial eligibility, and differential diagnosis.
-
Data Structure and Metadata: Each entry in BioInstruct follows a structured format containing three fields: a natural language instruction, an input argument, and the expected textual output. To organize the data, the authors used GPT-4 to classify instructions into four major categories: Question Answering (22.8%), Information Extraction (33.8%), Generation (33.5%), and Other tasks (10%). This classification was validated through human inter-rater reliability analysis, achieving a Krippendorff's alpha of 0.83.
-
Processing and Filtering Rules: To ensure high quality and diversity, the authors applied several filtering constraints during the generation process:
- Diversity: New instructions were only added to the pool if their ROUGE-L similarity to existing instructions was below 0.7.
- Keyword Filtering: Instructions containing terms like "image," "picture," or "graph" were removed as they are typically unsuitable for language models.
- Heuristic Filtering: Invalid generations were discarded if the instruction was too long (over 150 words), too short (under 3 words), or if the output was a repetition of the input.
-
Model Usage and Evaluation: The authors used BioInstruct for fine-tuning models to explore multi-task performance. They conducted ablation studies by training on specific subsets of tasks to determine their contribution to benchmark performance. For evaluation, they utilized several specialized biomedical benchmarks, including MedQA-USMLE, MedMCQA, PubMedQA, and BioASQ MCQA for question answering, as well as Conv2note and ICliniq for clinical generation tasks.
Method
The proposed methodology is structured around two fundamental stages: the construction of a specialized dataset named BioInstruct and the subsequent fine-tuning of various Large Language Models (LLMs).
The authors first focus on the development of the BioInstruct dataset, which serves as the foundational knowledge base for the model. This dataset is designed to bridge the gap between general linguistic capabilities and specialized biological expertise. By curating high-quality, domain-specific instructions, the authors ensure that the subsequent training process targets the nuances of biological reasoning and terminology.
Following the dataset preparation, the methodology transitions to the fine-tuning phase. During this stage, several LLM architectures are subjected to supervised fine-tuning using the BioInstruct corpus. This process aims to align the pre-trained models with the specific requirements of the biomedical domain, enhancing their ability to follow complex biological instructions and provide accurate, context-aware responses.
Experiment
The study evaluates the effectiveness of BioInstruct, a newly introduced automated instruction-tuning dataset, by fine-tuning LLaMA models across question answering, information extraction, and text generation tasks. The experiments validate that instruction tuning significantly enhances model performance in the biomedical domain, often outperforming established domain-specific baselines. Findings indicate that while increasing data volume generally improves capabilities, the specific combination of tasks that yields the best results varies depending on the target application.
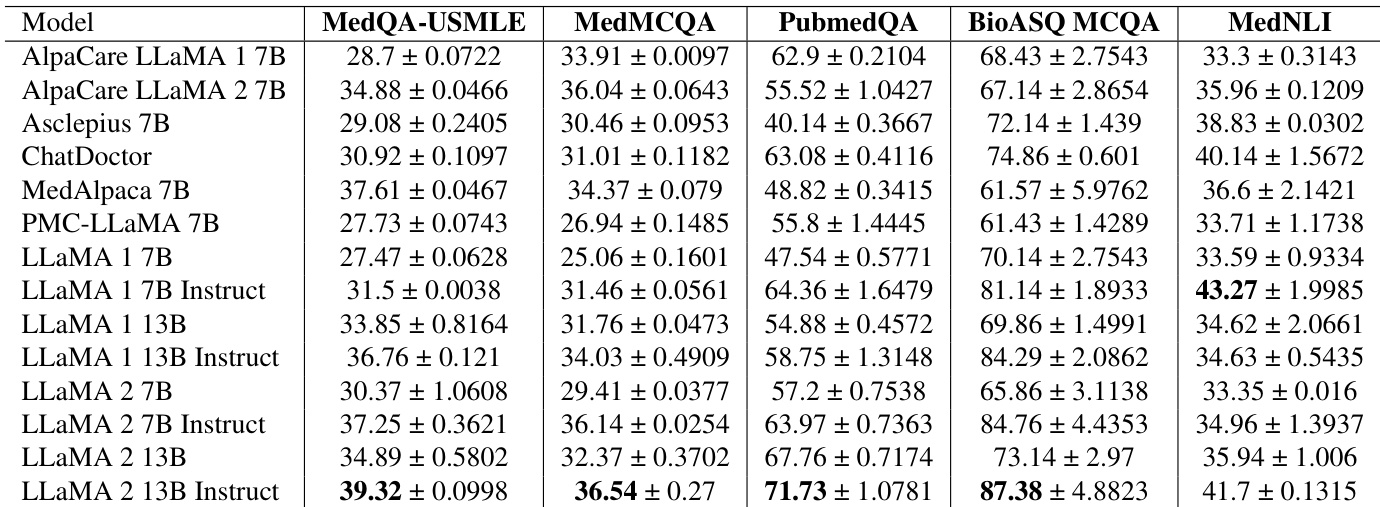
The authors evaluate several large language models on multiple-choice question answering and natural language inference tasks within the biomedical domain. Results show that instruction-tuned LLaMA models generally achieve higher performance across these benchmarks compared to their non-instructed versions and several specialized baseline models. Instruction-tuned LLaMA models demonstrate superior performance in medical question answering and inference tasks compared to base models The LLaMA 2 13B Instruct model achieves the highest scores across most evaluated biomedical question answering benchmarks Specialized baselines such as ChatDoctor and MedAlpaca show competitive results but are often surpassed by the instruction-tuned LLaMA variants
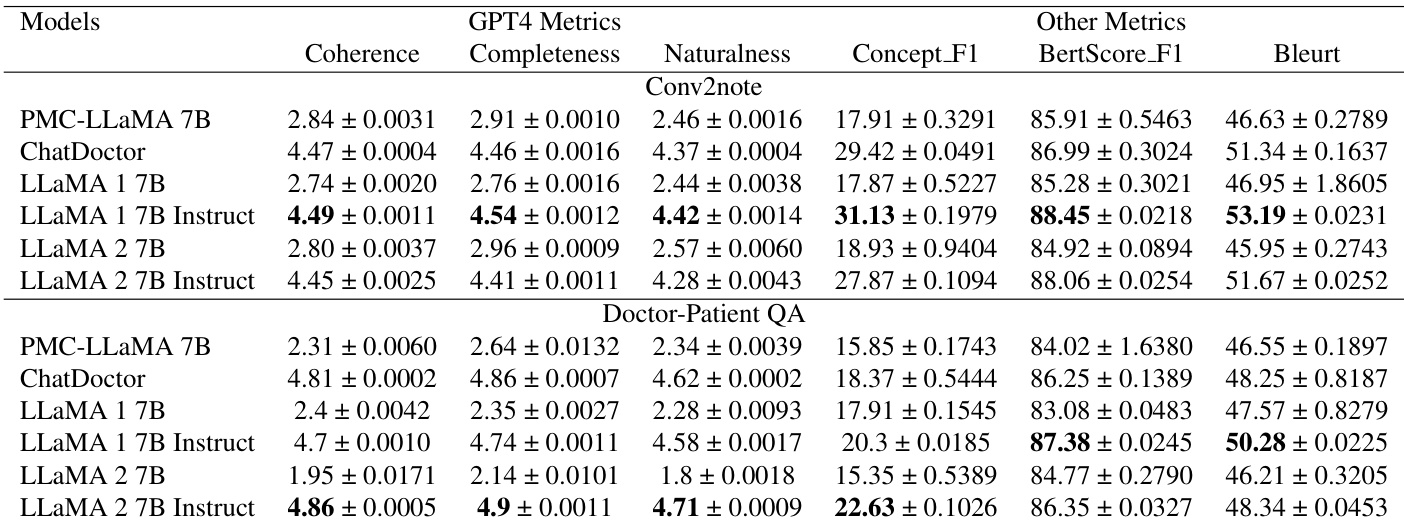
The authors evaluate several models on two text generation tasks, Conv2note and Doctor-Patient QA, using metrics such as GPT-4 based scores and semantic similarity measures. Results show that instruction-tuned LLaMA models generally achieve higher scores in coherence, completeness, and naturalness compared to their non-instructed versions and several baseline models. Instruction-tuned LLaMA 1 7B achieves superior performance in coherence and completeness for the Conv2note task compared to other models. For Doctor-Patient QA, the instruction-tuned LLaMA 2 7B model demonstrates higher coherence and naturalness scores than the baseline models. The instruction-tuned LLaMA 1 7B model shows improved semantic similarity and concept overlap in the Conv2note task relative to the original LLaMA 1 7B.
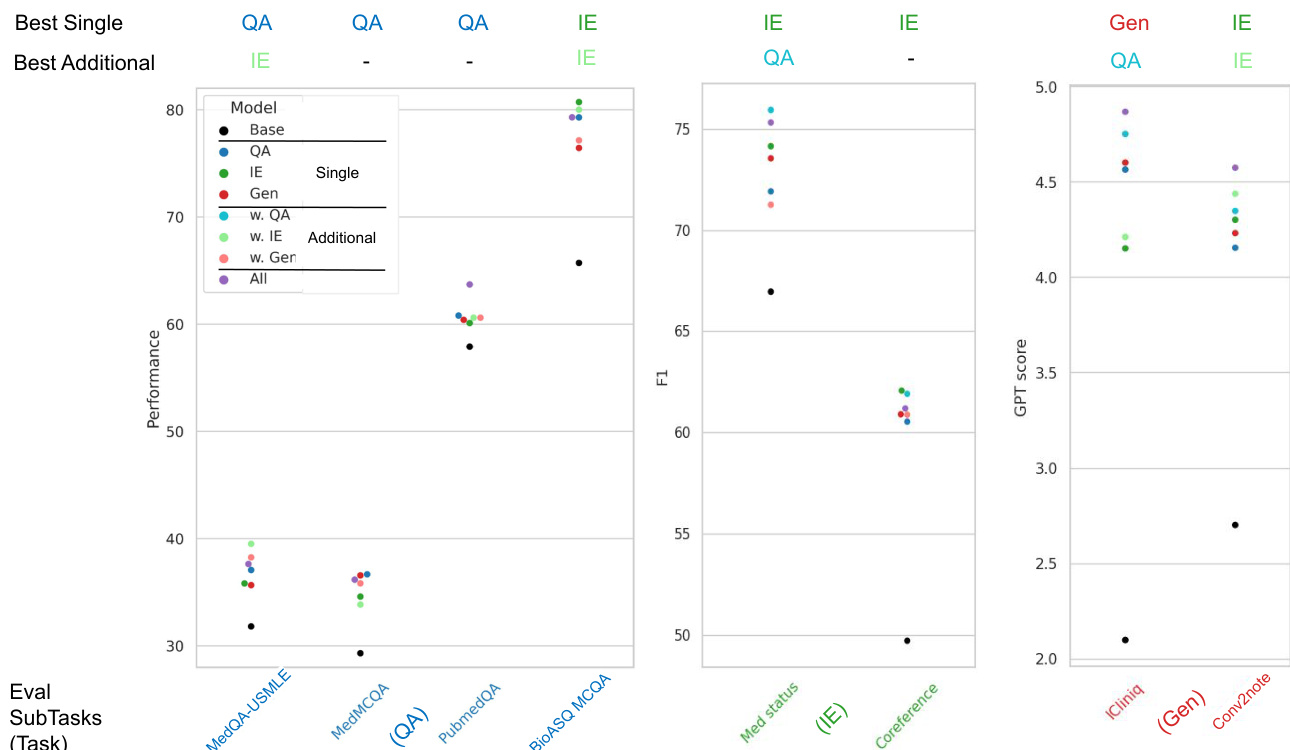
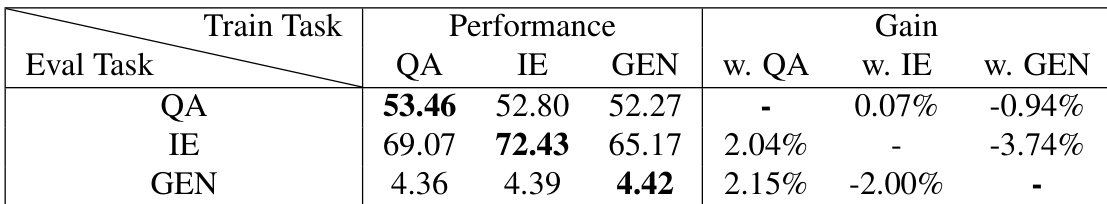
The authors evaluate the impact of multi-task instruction tuning by examining how training on specific task categories affects performance across different evaluation tasks. Results show that while certain task combinations provide synergistic benefits, the effectiveness of additional training tasks depends on the specific evaluation objective. Training with additional question answering tasks provides a performance boost for both information extraction and generation tasks. Information extraction tasks show a slight performance benefit when supplemented with question answering training. The performance gains from multi-task tuning are task-dependent, as some task combinations can lead to performance decreases in certain evaluation categories.
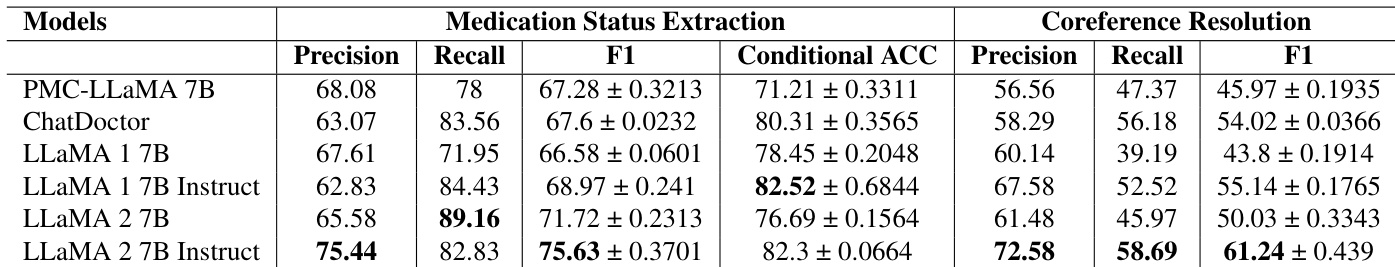
The the the table compares the performance of several biomedical language models on medication status extraction and coreference resolution tasks. Results demonstrate that instruction-tuned models generally achieve higher performance levels compared to their non-instructed counterparts and several domain-specific baselines. The instruction-tuned LLaMA 2 7B model achieves the highest precision and F1 scores for medication status extraction. For coreference resolution, the instruction-tuned LLaMA 2 7B model shows superior performance in precision, recall, and F1 scores compared to all other listed models. Instruction tuning provides notable improvements in recall for medication status extraction across different model versions.
The authors evaluate various LLaMA models across biomedical multiple-choice questions, natural language inference, text generation, and information extraction tasks to assess the impact of instruction tuning. The results demonstrate that instruction-tuned models consistently outperform their base versions and several specialized baselines in terms of coherence, naturalness, and extraction accuracy. Furthermore, while multi-task instruction tuning can provide synergistic benefits for information extraction and generation, the effectiveness of adding specific training tasks remains highly dependent on the target evaluation objective.