Command Palette
Search for a command to run...
From Computer Vision to Medical AI, Shanghai Jiao Tong University's Xie Weidi Published a Number of Results, Which Were Published in Nature sub-journals/NeurIPS/CVPR, etc.

In recent years, the development of AI for Science has accelerated, not only bringing innovative research ideas to the scientific research field, but also broadening the implementation channels of AI and providing it with more challenging application scenarios. In this process, more and more AI researchers have begun to pay attention to traditional scientific research fields such as medicine, materials, and biology, exploring the research difficulties and industry challenges therein.
Xie Weidi, a tenured associate professor at Shanghai Jiao Tong University, has been deeply engaged in the field of computer vision. He returned to China in 2022 and devoted himself to the research of medical artificial intelligence.At the COSCon'24 AI for Science forum co-produced by HyperAI, Professor Xie Weidi shared the team's achievements from multiple perspectives, including open source dataset construction and model development, under the title "Towards Developing Generalist Model For Healthcare".

HyperAI has organized and summarized the in-depth sharing without violating the original intention. The following is a transcript of the highlights of the speech.
Medical artificial intelligence is an inevitable trend
Medical research is of vital importance to everyone’s life and health. At the same time, the problem of uneven distribution of medical resources has not been fundamentally solved for a long time.So we hope to promote universal medical care and help everyone get high-quality diagnosis and treatment.
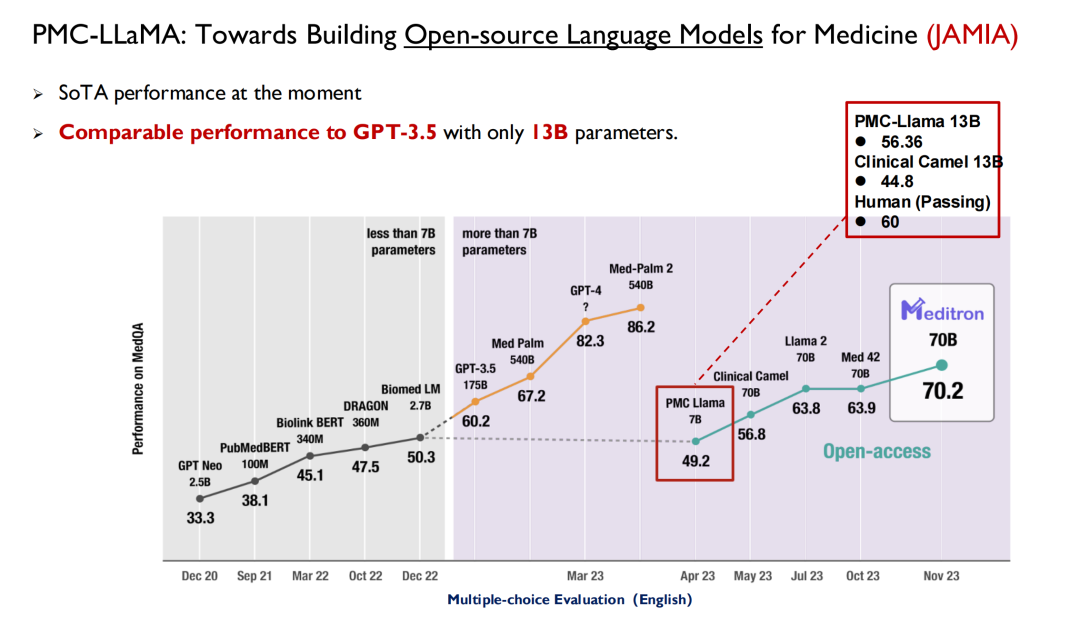
ChatGPT and other large models released in recent years have all used healthcare as the main battlefield for performance testing. As shown in the figure below, in the United States Medical Licensing Examination, large models will be able to reach a score of 50 before 2022, while humans will be able to reach 70, so AI has not attracted much attention from doctors.
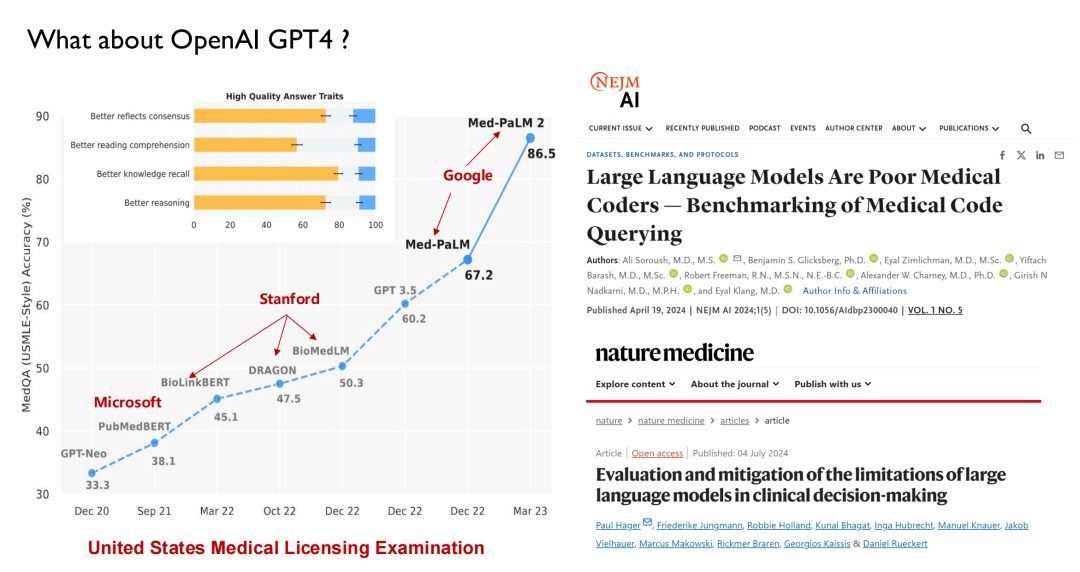
With the release of GPT 3.5, its score reached 60.2, a significant improvement. Later, Google released Med-PaLM and its updated version, with the highest score reaching 86.5. Today, GPT-4 can reach 90 points. Such high performance and iteration speed have made doctors begin to pay attention to AI.Many medical schools now offer a new discipline: intelligent medicine.
Likewise, not only medical students need to learn about artificial intelligence;AI students can also learn medical knowledge in their final year.Harvard University and other institutions have already set up relevant courses in AI majors.
But on the other hand, studies in academic journals such as Nature Medicine show thatThe big language model actually doesn’t understand medicine.For example, the big model currently does not understand ICD codes (diagnosis codes in the International Classification of Diseases) and it is difficult for it to provide timely guidance on the next step of medical treatment based on the patient's examination results like a doctor. It can be seen that the big model still has many limitations in the medical field.I think it will never replace doctors, and what our team wants to do is to make these models better assist doctors.
The team's primary goal: to build a general medical artificial intelligence system
I returned to China in 2022 and started to conduct research on medical artificial intelligence, so today I will mainly share the results of the team in the past two years. The medical industry covers a wide range, and the model we developed cannot be said to be universal, but we hope to cover as many important tasks as possible.
As shown in the figure below,On the input side, we hope to support multiple modes.For example, images, audio, patient health records, etc. After being input into the Multi-modal Generalist Model for Medicine, doctors can interact with it.The output of the model has at least two forms, one is visual,The location of the lesion is found through segmentation and detection.The second is text (Text),Output diagnosis results (Diagnosis) or reports (Report).
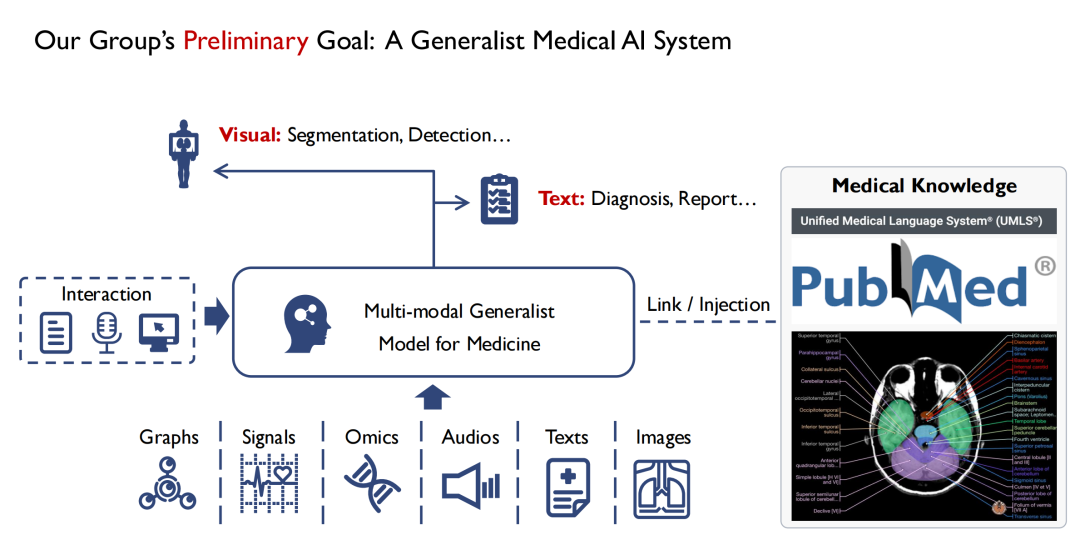
I am a computer vision major. From my observation, a big difference between vision and medicine is that most of the knowledge in medicine, especially evidence-based medicine, is summarized from human experience. If a beginner can exhaust all medical books, he can at least become a medical expert in theory. Therefore,During the model training process, we also hope to inject all medical knowledge into it.Because if the model lacks basic medical knowledge, it will be difficult to gain the trust of doctors and patients.
So, in summary,Our team’s primary goal is to build a multimodal universal medical model and inject as much medical knowledge into it as possible.
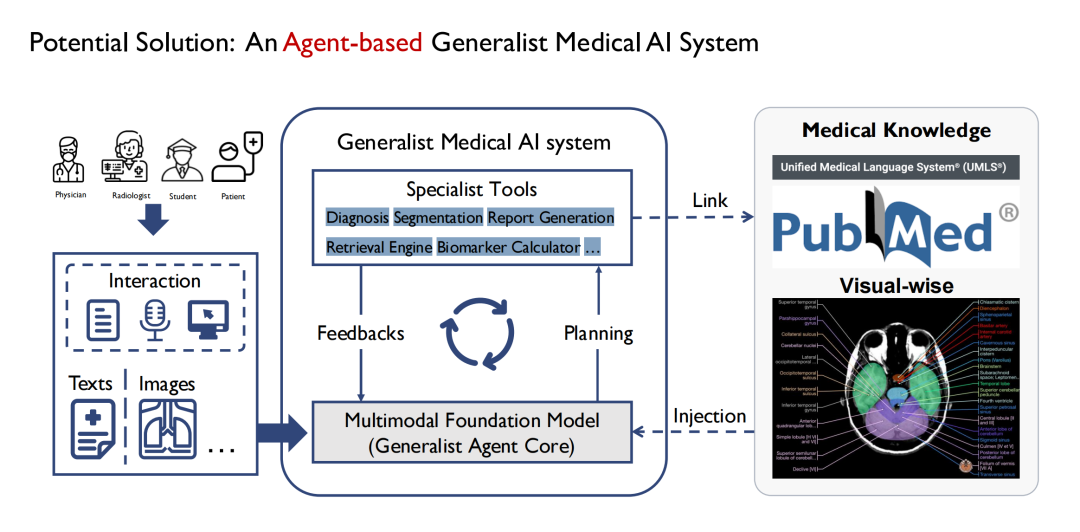
Initially, we started to define a general model, and gradually found that it was not realistic to build an omnipotent medical model like GPT-4. Because there are many departments in the hospital, and each department has different tasks, it is difficult for a general model to cover all tasks.So we choose to implement it through Agent.As shown in the figure below, the general model in the middle is composed of multiple sub-models, and each sub-model is essentially an Agent, and the general model is finally constructed in the form of Multi Agent.
Its advantages are that different agents can accept different inputs, so the input end of the model can be more complex and diversified; multiple agents can also form a chain of thinking in the process of handling different tasks step by step; the output end is also richer, for example, one agent can complete the segmentation of multiple types of medical images such as CT and MRI; at the same time, it also has better scalability.
Contribute high-quality open source datasets
Focusing on the big goal of building a multimodal universal medical model, I will now introduce the team's achievements from multiple aspects, including open source datasets, large language models, disease diagnosis agents, etc.
The first is our contribution to open source datasets.
There is no shortage of data sets in the medical field, but due to privacy issues, open and available high-quality data is relatively scarce. As an academic team, we hope to contribute more high-quality open source data to the industry.So after I returned to China, I started to build a large-scale medical dataset.
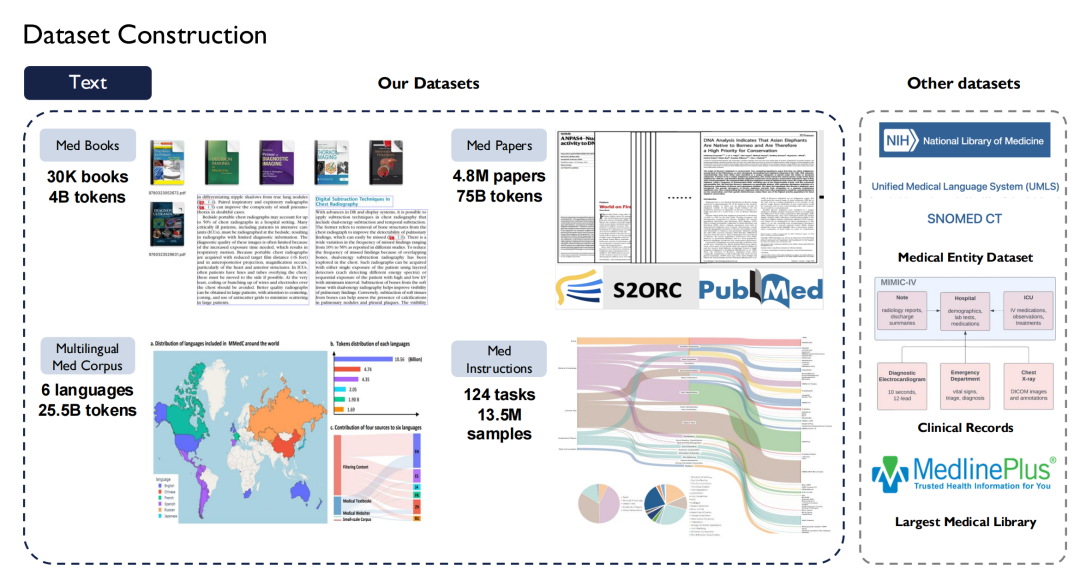
In terms of text, we collected more than 30,000 medical books, containing 4 billion tokens; crawled all medical literature in PubMed Central (PMC), including 4.8 million papers and 75 billion tokens; and collected medical books in eight languages including Chinese, English, Russian, and Japanese on the Internet and converted them into text.
also,We also built Super Instructions in the medical field.Taking into account the diversity of tasks, 124 medical tasks are listed, involving 13.5 million samples.

Text data is relatively easy to obtain, but vision-language (image-text pairs) is more difficult to obtain. We crawled about 200,000 cases from the Radiopaedia website, collected images and their captions in papers, and obtained more than 30,000 volumes from basic reports of radiology departments.
Currently, most of our data is open source.
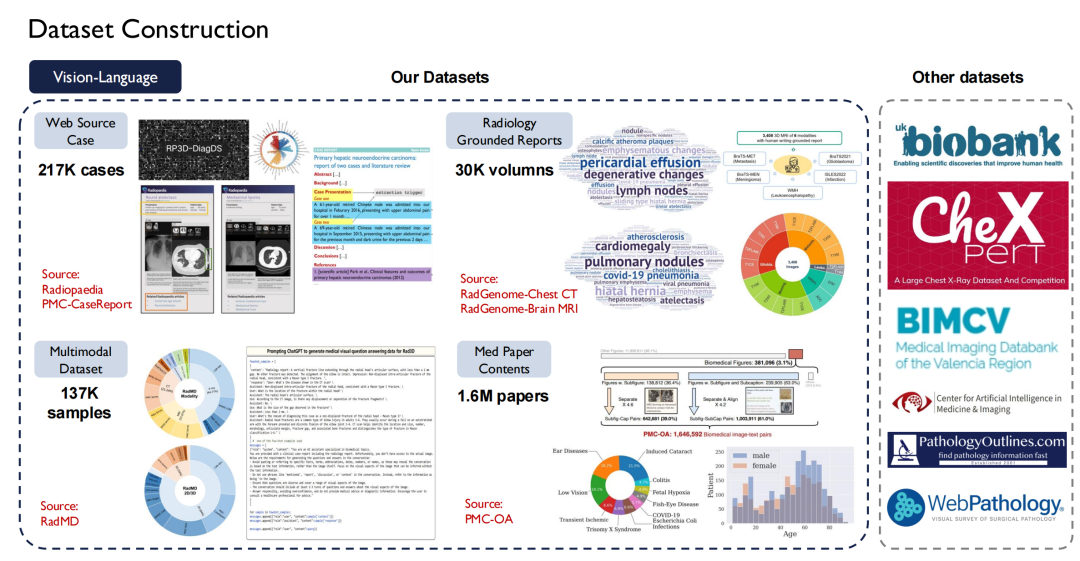
The right side of the above figure shows other public data sets, such as UK Biobank, for which we paid to purchase data on nearly 100,000 patients in the UK for 10 years; in addition, Pathology Outlines provides comprehensive pathology knowledge.
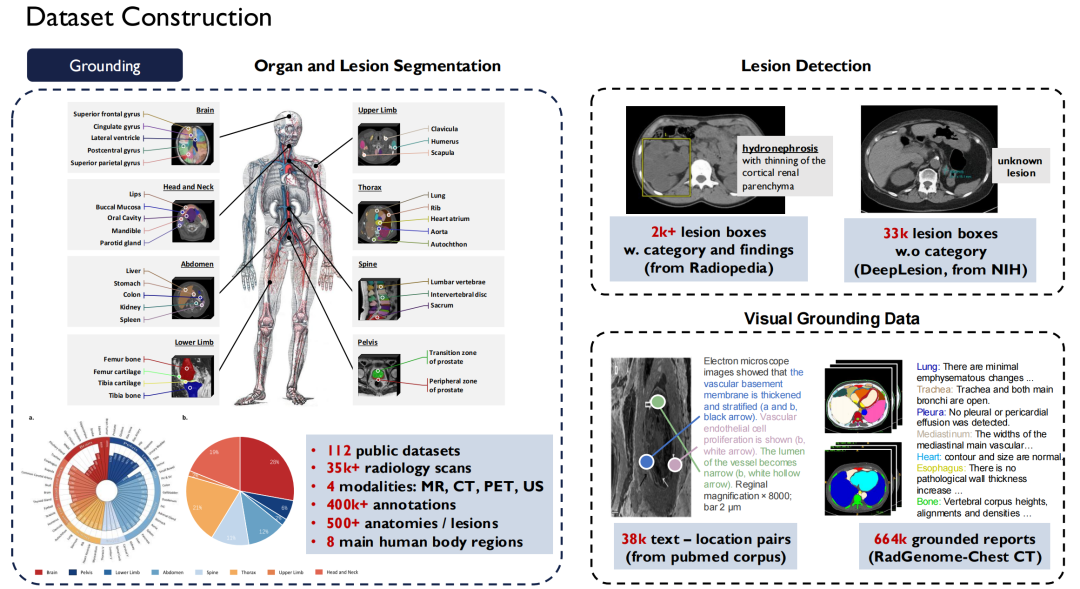
In terms of Grounding Data,This is the segmentation and detection data I just mentioned.We unified nearly 120 radiology image public datasets available on the market into one standard, resulting in more than 35,000 2D/3D radiology scan images.It covers four modalities: MR, CT, PET, and US, with 400,000 fine-grained annotations, and these data cover 500 organs in the body.At the same time, we have expanded the description of lesions and made all these data sets open source.
Continuous iteration to create a professional medical model
Language Model
Only high-quality open source datasets can help students and researchers to better train models. Next, I will introduce the team's achievements in the model.
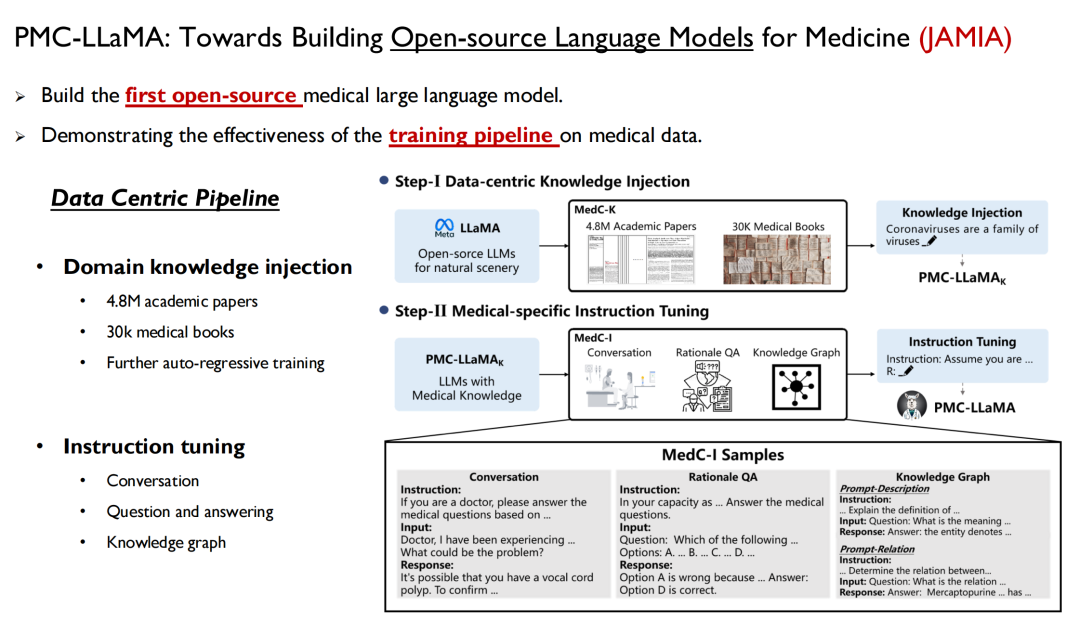
The first is the language model, which is a way to quickly inject human knowledge into the model. Last April, we launched a model called PMC-LLaMA, and the related research was published in JAMIA under the title "Towards Building Open-source Language Models for Medicine".
Paper address:
https://academic.oup.com/jamia/article/31/9/1833/7645318
This is the first open source large language model we have developed in the medical field. We trained all the medical data and the paper data mentioned above into the model, performed autoregressive training, and then fine-tuned the instructions to convert the data into question-answer pairs.
Yale University researchers mentioned in their paper: PMC-LLaMA is the earliest open source medical model in the field.Many researchers later used it as a baseline, but in my opinion, there is still a gap between PMC-LLaMA and closed-source models, so we will continue to iterate and upgrade this model in the future.
Subsequently, we published another paper in Nature Communications: "Towards Building Multilingual Language Models for Medicine".We have launched a multilingual medical model that covers six languages: English, Chinese, Japanese, French, Russian, and Spanish, and is trained with 25 billion medical-related tokens. Due to the lack of a unified multilingual standard test set, we have also built a related benchmark for everyone to test.
In practice, we have found that as the base model is upgraded and medical knowledge is injected into it, the performance of the resulting large medical model will also be improved.
Most of the tasks mentioned above are "multiple-choice questions", but we all know that in the actual work of doctors, it is impossible to only do multiple-choice questions, so we hope that the large language model can be embedded in the doctor's workflow in the form of free text.In our new research, we will focus more on clinical tasks, collect relevant data sets, and improve the clinical scalability of the model.
The relevant paper is still under review.
Visual-language Model
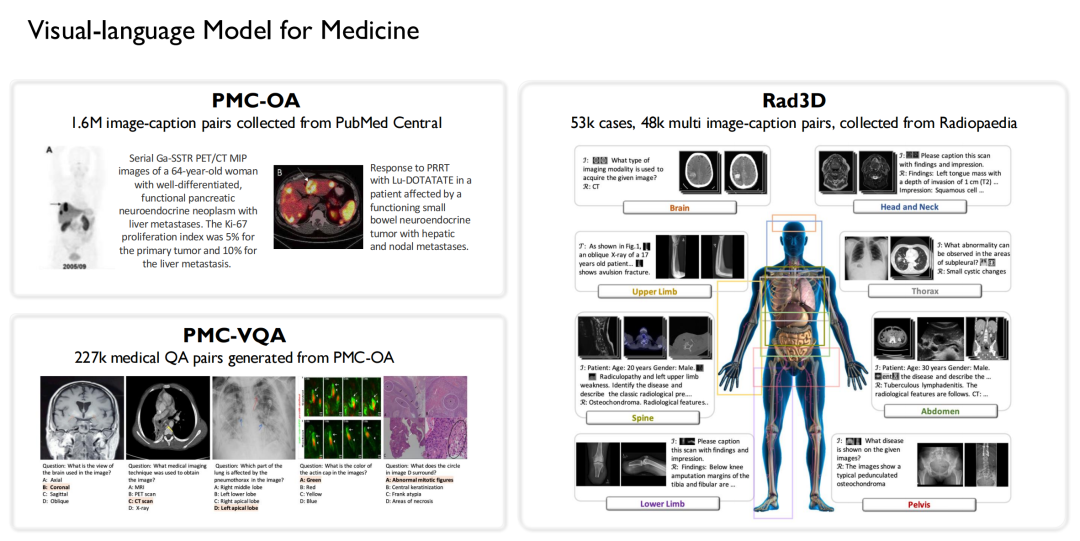
Similarly, we are also one of the earliest teams to conduct research on visual-language models in the medical field. Based on the data mentioned above,We built 3 open source datasets:
* Collected 1.6 million large-scale image-caption pairs from PubMed Central and constructed the PMC-OA dataset;
* Generated 227,000 medical visual question-answer pairs from PMC-OA to form PMC-VQA;
* A Rad3D dataset was constructed by collecting 53,000 cases and 48,000 multiple image-caption pairs from the Radiopaedia species.
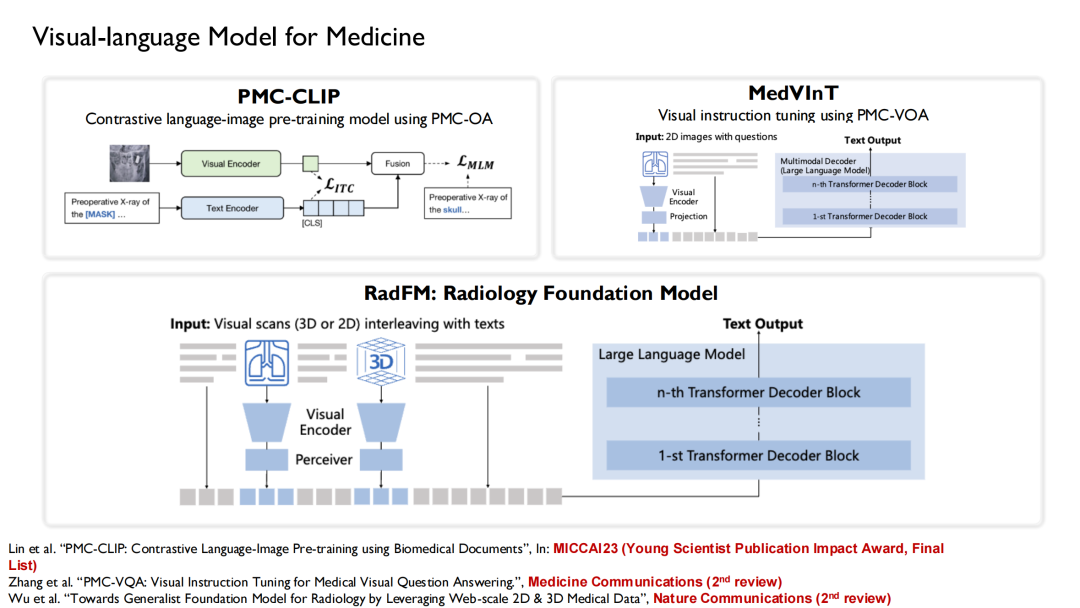
Based on these data sets, we combined the language models that have been trained.Three versions of vision-language models were trained: PMC-CLIP, MedVInT, and RadFM.
PMC-CLIP is a result we published at MICCAI 2023, the top conference in the field of medical artificial intelligence imaging.Finally, he was selected as the "Young Scientist Publication Impact Award, Final List".The award is given to 3 to 7 winning papers selected from the papers published in the past five years.
RadFM (Radiology Foundation Model) is now quite popular, and many researchers use it as a baseline. During the training process,We input text-image interweaving into the model, which can directly generate answers based on questions.
Enhance domain-specific knowledge and improve model performance
The so-called knowledge-enhanced representation learning (Knowledge-enhanced Representation Learning) needs to solve the problem of how to inject medical knowledge into the model. We have also conducted a series of research around this challenge.
First of all, we need to solve the problem of where "knowledge" comes from.On the one hand, it is General Medical Knowledge.Sourced from the Internet, as well as relevant papers and books sold by UMLS, the largest knowledge graph in the medical field;On the other hand, there is domain-specific knowledge.For example, case reports, radiological images, ultrasound, etc.; as well as anatomical knowledge, they can all be obtained from some websites. Of course, special attention should be paid to copyright issues here, as the content on some websites cannot be used.

After obtaining this "knowledge", we can draw a knowledge graph.This establishes relationships between disease and disease, drug and drug, and protein and protein, and provides detailed descriptions.
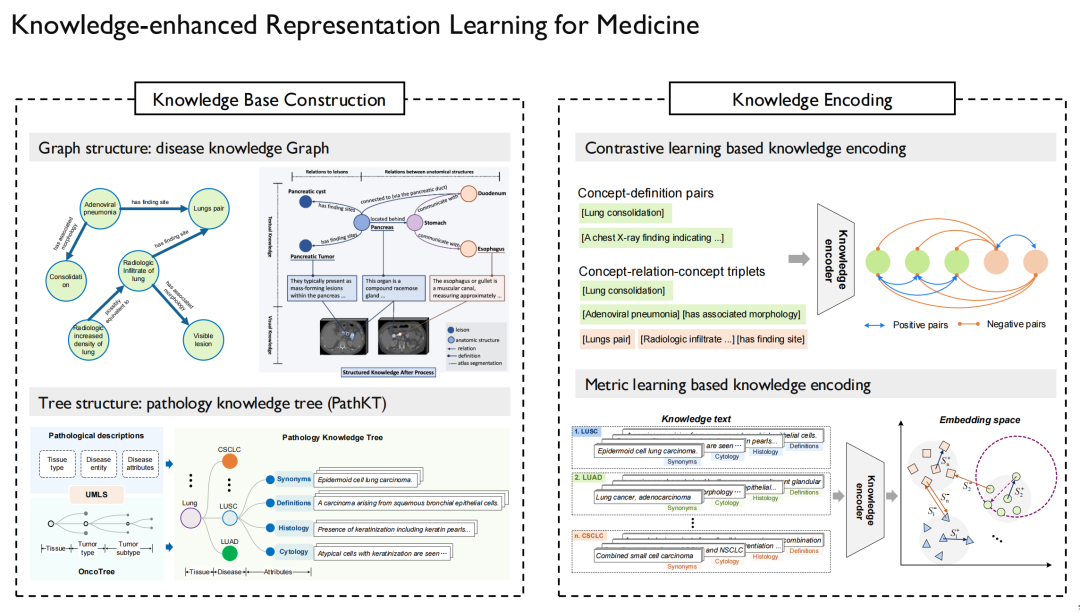
On the left side of the above picture is the pathology knowledge graph and knowledge tree we built.It is mainly aimed at cancer diagnosis, because cancer may occur in various organs of the human body and is also divided into different subtypes, which is suitable for making a tree structure. Similarly, in addition to multimodal pathology, we have also conducted related research on multimodal radiology and multimodal X-ray.
The next step is to inject this knowledge into the language model so that the model can remember the relationship between the graph and the points in the graph.Once the language model is trained, the visual model only needs to be aligned to the language model.
We compared our results with those of Microsoft and Stanford, and the results showed thatThe model with added domain knowledge has much higher performance than other models without domain knowledge.
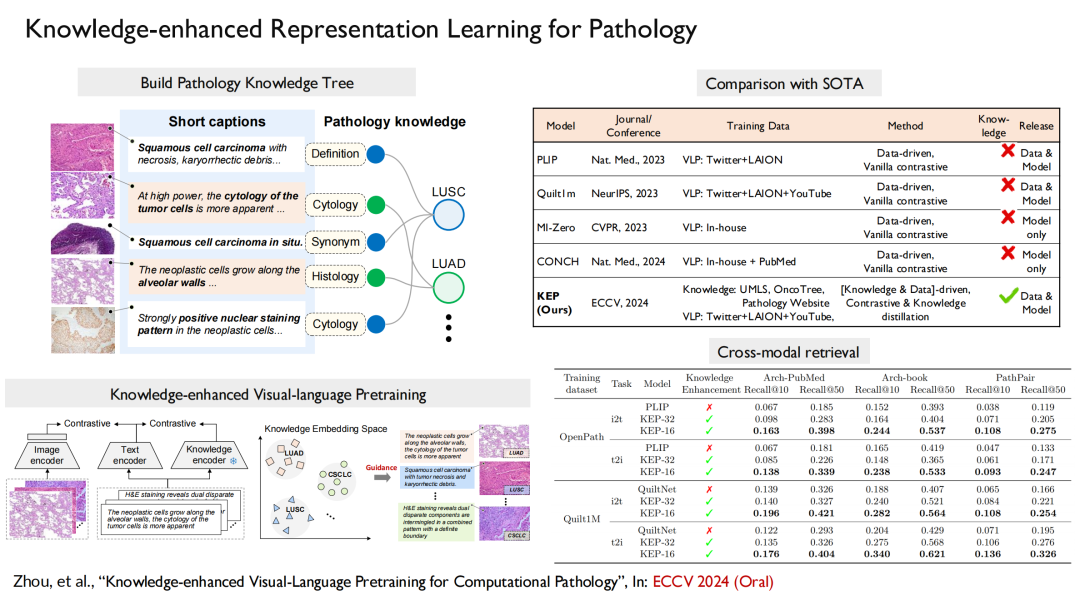
For pathology, our paper "Knowledge-enhanced Visual-Language Pretraining for Computational Pathology" was selected for the top machine learning conference ECCV 2024 (Oral). In this achievement, we built a knowledge tree, injected it into the model training, and then aligned vision with language.
Furthermore, we used the same method to construct a multimodal radiology imaging model, and the results were published in Nature Communications under the title "Large-scale long-tailed disease diagnosis on radiology images".The model can directly output the corresponding symptoms based on the patient's radiological images.
In summary,Our work implemented a complete process - first, we built the largest open source dataset of radiological images, which contains 200,000 images, 41,000 patient images, covering 930 diseases, etc.; second, we built a multimodal, multilingual model to enhance knowledge in specific fields; finally, we built a corresponding benchmark.
About Professor Xie Weidi
He is a tenured associate professor at Shanghai Jiao Tong University, a recipient of the National (Overseas) High-level Young Talent Program, Shanghai Overseas High-level Talent Program, and Shanghai Morning Star Program. He is the young project leader of the Ministry of Science and Technology's Science and Technology Innovation 2030 - "New Generation Artificial Intelligence" major project, and the project leader of the National Natural Science Foundation of China.
He received his Ph.D. from the Visual Geometry Group (VGG) at the University of Oxford, where he studied under Professor Andrew Zisserman and Professor Alison Noble. He is one of the first recipients of the Google-DeepMind full scholarship, the China-Oxford Scholarship, and the Oxford University Engineering Department Outstanding Award.
His main research areas are computer vision and medical artificial intelligence. He has published more than 60 papers, including CVPR, ICCV, NeurIPS, ICML, IJCV, Nature Communications, etc., with more than 12,500 citations on Google Scholar. He has won the Best Paper Award, Best Poster Award, and Best Journal Paper Award at top international conferences and seminars many times, and is a MICCAI Young Scientist Publication Impact Award Finalist. He is a special reviewer for Nature Medicine and Nature Communications, and is the field chair of CVPR, NeurIPS, and ECCV, the flagship conferences in the field of computer vision and artificial intelligence.
* Personal homepage:










