Command Palette
Search for a command to run...
First! Four Universities Jointly Launched the Drug Development Language Model Y-Mol, Which Is Ahead of LLaMA2 in Performance

Large language models represented by ChatGPT, ChatGLM and LLaMA have become powerful tools for people to explore the unknown world. These models with billions of parameters have demonstrated strong capabilities in generating text and understanding context through careful training of large-scale text corpora. However, most of these models perform well in general tasks, but face considerable challenges in certain specific fields, especially drug research and development.
Unlike the field of natural language processing, the field of drug development lacks a unified standard paradigm, and the development process is complex and costly. In addition, it involves multiple disciplines such as computational chemistry, structural biology, and bioinformatics. Relevant data is difficult to obtain, and the interaction data between drug-related entities requires sophisticated domain knowledge to be labeled.These factors together limit the application of large language models in drug research and development.
In response to this, research teams from Hunan University, Central South University, Hunan Normal University, and Xiangtan University jointly proposed a large language model Y-Mol guided by multi-scale biomedical knowledge. Y-Mol is an autoregressive sequence-to-sequence model that can be fine-tuned on different text corpora and instructions, greatly enhancing the performance and potential of the model in drug development. This is a new breakthrough in the field of drug development using large language models.
The study, titled "Y-Mol: A Multiscale Biomedical Knowledge-Guided Large Language Model for Drug Development", has been published as a preprint on arxiv.
Research highlights:
* Y-Mol is the first large language model built for drug discovery
* Y-Mol builds an information-rich instruction dataset by integrating multi-scale biomedical knowledge
* Y-Mol excels in drug-drug interactions, drug-target interactions, molecular property prediction, and demonstrates strong capabilities in understanding and versatility in various drug development tasks

Paper address:
https://doi.org/10.48550/arXiv.2410.11550
Follow the official account and reply "drug development model" to get the complete PDF
The open source project "awesome-ai4s" brings together more than 100 AI4S paper interpretations and provides massive data sets and tools:
https://github.com/hyperai/awesome-ai4s
Fully explore two types of data sets to build a comprehensive biomedical corpus
In terms of constructing the pre-training dataset for Y-Mol, the study selected two types of datasets:A text corpus from biomedical PubMed publications; supervised instructions built on a biomedical knowledge graph, and inference data extracted from expert models.
In order to deeply explore the rich biomedical knowledge in publications,The study extracted and pre-processed more than 33 million publications covering multiple disciplines from online publishing platforms such as PubMed.As shown in Figure A below, the researchers extracted visible abstracts and introductions from these publications as biomedical text data (Reconstructed Text) to ensure the quality and relevance of the corpus.
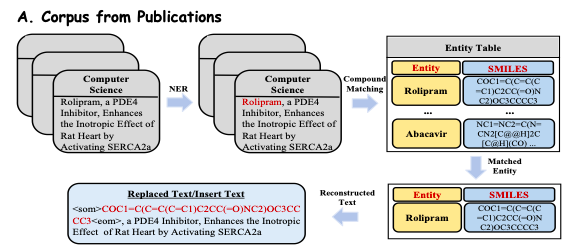
In order to efficiently extract domain knowledge from the biomedical knowledge base, this study converts the facts in the knowledge base into natural language prompts.As shown in Figure B below, the study considers that each reasoning chain in the subgraph has clear relational semantics, so each coherent path is extracted and converted into a natural language description using a carefully designed template as a prompt context. The study then combines these constructed contexts with the corresponding questions and inputs them into Y-Mol to output supervised answers.
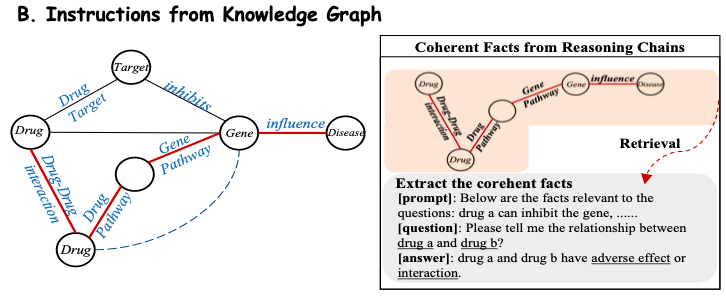
In addition, in order to obtain large-scale instructions based on drug attributes and domain knowledge, this study used expert synthetic data from existing small models to construct instructions and refined the drug knowledge spectrum into Y-Mol.Ultimately, the study assembled 11.2 million corpus entries and 2.3 million carefully crafted instructions.
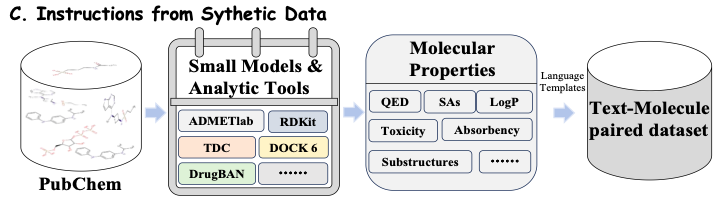
As shown in Figure C below, for a given drug molecule, in order to extract more comprehensive molecular properties (Molecular Properties), this study brings together a series of advanced molecular tools and computational models, such as ADMETlab, RDKit, TDC and DrugBAN. These tools and models extract molecular information with different characteristics from publicly available data sets, including QED, SAs, LogP, toxicity (Toxicity), absorbency (Absorbency) and substructures (Substructures). In this way, the study can continuously integrate the latest models and tools and use their prediction data to train the model, so that Y-Mol can evolve in real time and maintain its leading position in the field of drug research and development.
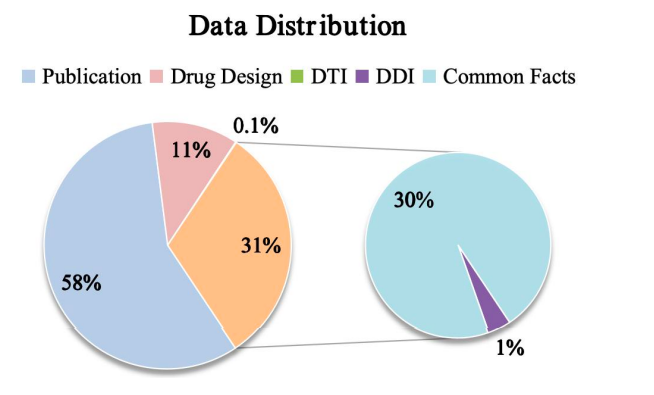
Finally, as shown in the figure below, the study shows the data distribution of Y-Mol for different tasks during the pre-training and supervised fine-tuning stages. In terms of the evaluation of reasoning ability, in order to comprehensively test the performance of Y-Mol in drug-target interaction (DTI) prediction and drug-drug interaction (DDI) prediction,The research team selected the industry-wide recognized benchmark datasets DrugBank and DrugCentral for DTI prediction.
At the same time, in order to evaluate the performance of DDI prediction, the researchers used the dataset provided by Ryu and Deng.These evaluation methods were carefully selected to ensure that Y-Mol can be fairly and comprehensively tested under industry standards in the field of drug development to prove its effectiveness.
Ryu's dataset: https://doi.org/10.1073/pnas.1803294115
Deng's dataset: https://doi.org/10.1093/bioinformatics/btaa501
Y-Mol: Based on LLaMA2-7b, dedicated to drug development
This study selected LLaMA2-7b as the basic large language model to build an advanced training and reasoning framework specifically for drug development - Y-Mol. As shown in the figure below,The development of Y-Mol was divided into two key phases:
first,Y-Mol is pre-trained on a large-scale corpus of biomedical publications and fine-tunes LLaMA2 through self-supervised pretraining, enabling Y-Mol to have a basic grasp of the background knowledge of drug development.then,LLaMA2 was further supervised and fine-tuned using drug-related domain knowledge and expert synthetic data. This process input a large amount of drug-related information into Y-Mol, enhancing the model's understanding of the interaction mechanisms in the drug development process.
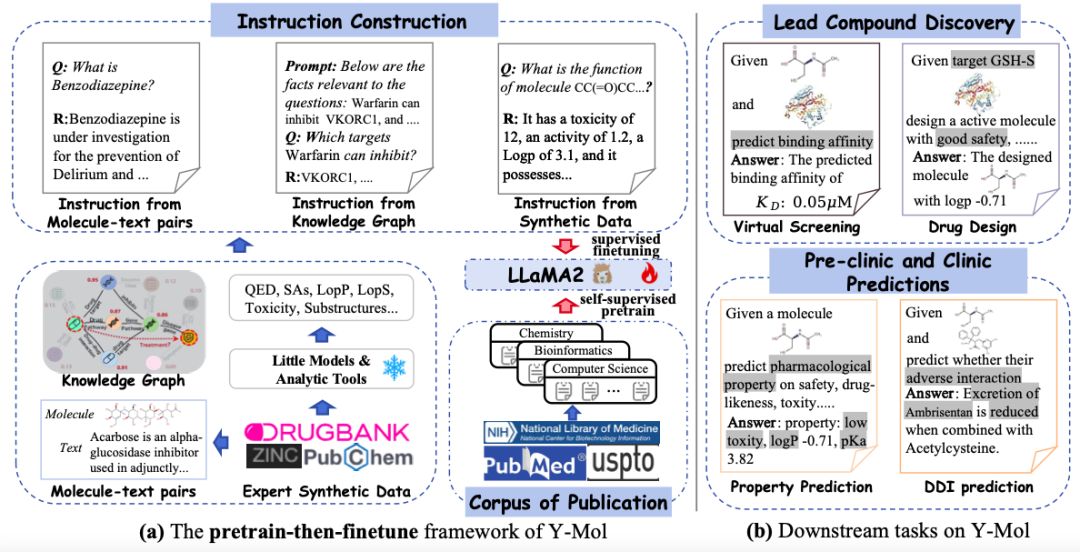
The study carefully designed a diverse set of instructions and fine-tuned Y-Mol. These instructions include instructions from Molecule-text Pairs and descriptions extracted from drug databases. These descriptions present the properties, structure and function of drugs in natural language and contain rich semantic information, which helps to enhance the consistency between humans and large language models in the perception of drug entities.
As shown in the figure below, the study uses the generated instructions as the input for supervised learning and feeds them into Y-Mol. Specifically, the constructed prompt contexts and questions are input into Y-Mol, and these constructed answers are used to supervise the output generated by the model.
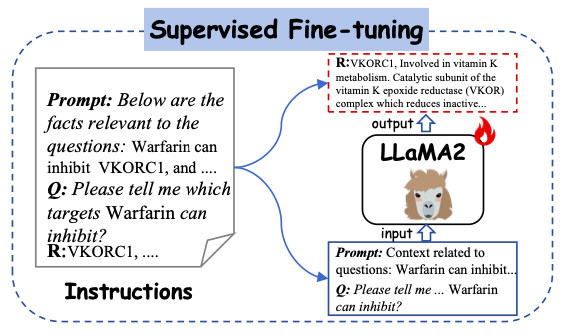
After carefully fine-tuning Y-Mol according to these generated instructions, the researchers applied it to a range of downstream tasks, covering multiple links from lead compound discovery to preclinical and clinical predictions. Through this supervised fine-tuning method, Y-Mol can more accurately understand and handle complex problems in drug development, providing a powerful tool for computer-aided drug development.
Research results: Y-Mol has the best prediction performance
In order to fully verify the effectiveness of Y-Mol in the field of drug research and development, the study carefully designed a series of tasks covering different stages such as lead compound discovery, pre-clinical research and clinical predictions.Specifically, the different key tasks are as follows: (1) virtual screening and drug design for lead compound discovery; (2) prediction of the physical and chemical properties of discovered lead compounds in the preclinical stage; and (3) prediction of potential adverse drug events in the clinical stage.
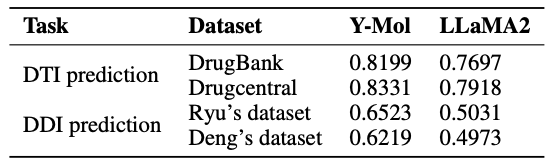
In virtual screening,Identifying unknown drug-target interactions is crucial. As shown in the table below, compared with LLaMA2, Y-Mol's AUC scores on DrugBank and DrugCentral datasets increased by 5.02% and 4.13%, respectively. This shows that Y-Mol performs well in DTI prediction of multi-scale data sources, demonstrating its superior performance in virtual screening.
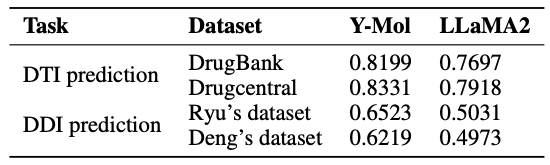
In drug design,In order to verify the performance of Y-Mol in discovering new lead compounds, the study also designed a task to produce effective compounds under specific conditions. That is, given a target condition and a descriptive query, it evaluates whether Y-Mol can accurately generate the corresponding SMILES sequence molecules from the context information.
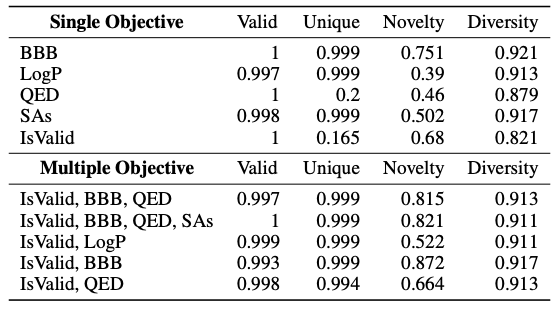
As shown in the table below, this study introduced standard indicators such as Valid, Unique, Novelty, and Diversity to predict different single objectives such as BBB and LogP. The results show that Y-Mol has better overall performance. In contrast, only the domain adaptability of the LLaMA2-7b model is poor and cannot generate effective molecules. At the same time, the study also tested the drug design performance of Y-Mol under multiple objectives. The results show that Y-Mol also performs well in this case.
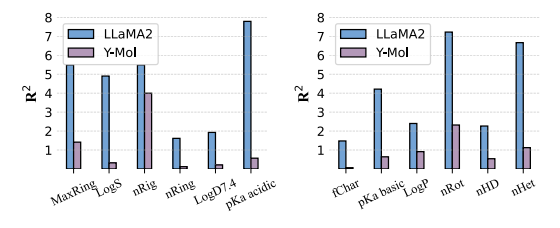
In molecular property prediction,As shown in the figure below, Y-Mol exhibits lower R² scores than LLaMA2 in all tasks, which indicates that Y-Mol has stronger generalization ability in predicting physicochemical properties.
During the clinical stage of drug development, predicting potential drug-drug interactions is key to ensuring safe drug use.As shown in the figure below, Y-Mol performs well in the task of identifying potential drug interactions (DDIs).
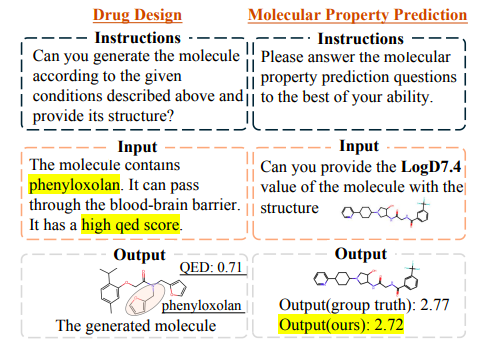
As shown in the figure below,The drugs designed by Y-Mol effectively meet the constraints set in the query. Similarly, Y-Mol can accurately predict the LogD7.4 of a given molecule, and the predicted results are very close to the actual values.This demonstrates the effectiveness of Y-Mol in solving drug development tasks.
AI technology: a new engine in drug development
In fact, in the long journey of drug development, scientists have been looking for new technologies that can accelerate the process. In recent years, AI technology has shown great application potential in this field. They can not only deeply understand the disease mechanism, but also play an important role in key stages such as drug discovery and clinical trials.
In the business world,Some companies have achieved remarkable results in AI drug development. For example, AI drug development company Insilico Medicine announced earlier this year that they had discovered a new clinical candidate drug for the treatment of idiopathic pulmonary fibrosis with a completely new mechanism, which has been verified by multiple human cell and animal model experiments. In addition, Huawei Cloud has cooperated with the Shanghai Institute of Materia Medica, Chinese Academy of Sciences, to launch the Pangu Drug Molecular Model, which can realize AI-assisted drug design for the entire process of small molecule drugs and improve the efficiency and accuracy of drug development.
In the field of scientific research,One of the authors of this study, Professor Zeng Xiangxiang's team at Hunan University, also designed a large language model for peptide sequences, and trained the model by gradually adding calculation and screening conditions. In just three months, the model successfully designed and synthesized 29 potential antimicrobial peptides, 26 of which showed broad-spectrum antimicrobial activity. In mouse experiments, three antimicrobial peptides showed antibacterial effects comparable to FDA-approved antibiotics, and no obvious drug resistance was observed during 25 days of continuous culture and monitoring. This achievement has been formally accepted by Nature Communications.
Paper link:
https://www.nature.com/articles/s41467-024-51933-2
In addition, another author of this study, Professor Cao Dongsheng of Central South University, together with Professor Hou Tingjun and Professor Xie Changyu of Zhejiang University, recently developed the molecular optimization tool Prompt-MolOpt. This algorithm uses the training strategy of prompt learning to realize the application of zero-shot learning and few-shot learning in multi-property optimization.
Paper link:
https://www.nature.com/articles/s42256-024-00916-5
From in-depth understanding of disease mechanisms to accelerating drug discovery and optimizing clinical trial design, AI technology is becoming a new engine for drug research and development. As technology continues to advance, it will play an increasingly critical role in future medical research.