Command Palette
Search for a command to run...
20 Experimental Data Create AI Protein Milestone! Shanghai Jiaotong University and Shanghai AI Lab Jointly Released FSFP to Effectively Optimize Protein pre-training Models

Proteins, these tiny but powerful biomolecules, are the basis of life activities and play a variety of roles in organisms. However, it is a very challenging task to precisely adjust and optimize protein functions to suit specific industrial or medical needs. Traditionally, scientists have relied on wet lab methods to explore the mysteries of proteins, but this method is time-consuming and expensive.
Fortunately, with the rapid development of artificial intelligence, a new tool, pre-trained protein language models (PLMs), is helping us understand and predict the behavior of proteins in an unprecedented way. PLMs learn the distribution characteristics of amino acid sequences in millions of proteins in an unsupervised manner, showing great potential in revealing the implicit relationship between protein sequences and their functions, thus helping to efficiently explore a large amount of design space. Today,Pre-trained PLMs have made significant progress in the absence of experimental data, but their accuracy and interpretability still need to be improved.In addition, traditional supervised learning models require a large number of labeled training samples, which is also an obstacle that is difficult to overcome in practical applications.
In order to solve the above problems,Professor Hong Liang's research group from the School of Natural Sciences/School of Physics and Astronomy/Zhangjiang Institute for Advanced Studies/School of Pharmacy of Shanghai Jiao Tong University, in collaboration with Tan Pan, a young researcher at the Shanghai Artificial Intelligence Laboratory,By combining meta-transfer learning (MTL), learning to rank (LTR), and parameter-efficient fine-tuning (PEFT),We developed a training strategy, FSFP, that can effectively optimize protein language models when data is extremely scarce.It can be used for small-sample learning of protein adaptability. It greatly improves the effect of traditional protein pre-training large models in mutation-property prediction when using very little wet experimental data, and also shows great potential in practical applications.
The related research was published in Nature Communications, a Nature subsidiary, under the title "Enhancing efficiency of protein language models with minimal wet-lab data through few-shot learning".

Paper address:
https://doi.org/10.1038/s41467-024-49798-6
ProteinGym protein mutation dataset download address:
The open source project "awesome-ai4s" brings together more than 100 AI4S paper interpretations and provides massive data sets and tools:
https://github.com/hyperai/awesome-ai4s
FSFP optimizes protein language model to address data shortage problem
The FSFP approach consists of three phases:Build auxiliary tasks for meta-training, Meta-train PLMs on the auxiliary tasks, and Transfer PLMs to the target task via LTR.
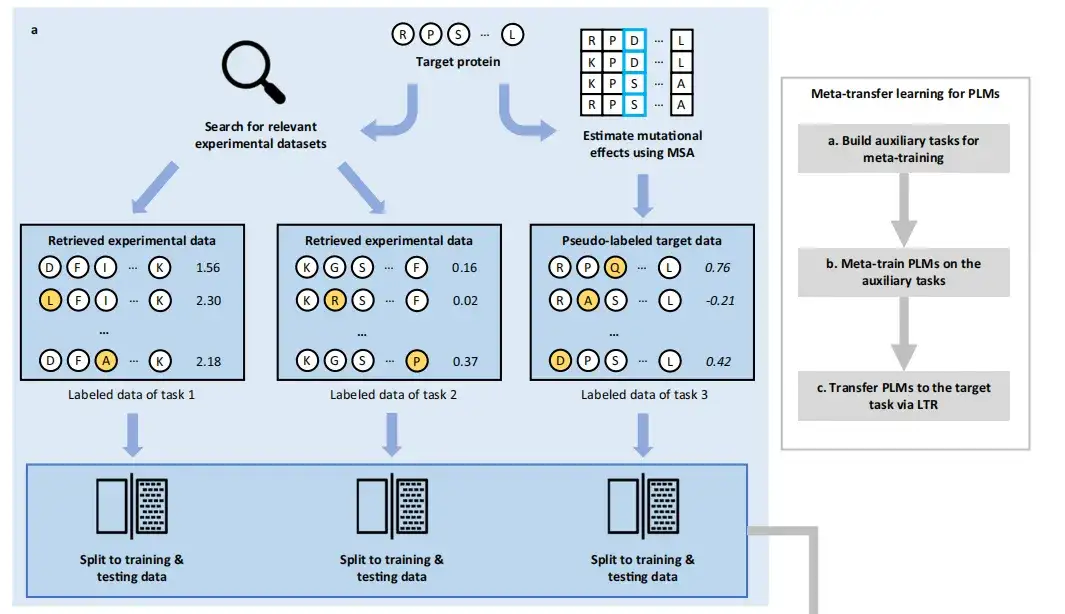
in,Meta-learning aims to accumulate experience from multiple learning tasks to train a model that can quickly adapt to new tasks using only a small number of training examples and iterations.Therefore, this study first used PLMs to encode the wild-type sequence or structure of the target protein and the sequence or structure in the database into an embedded vector.
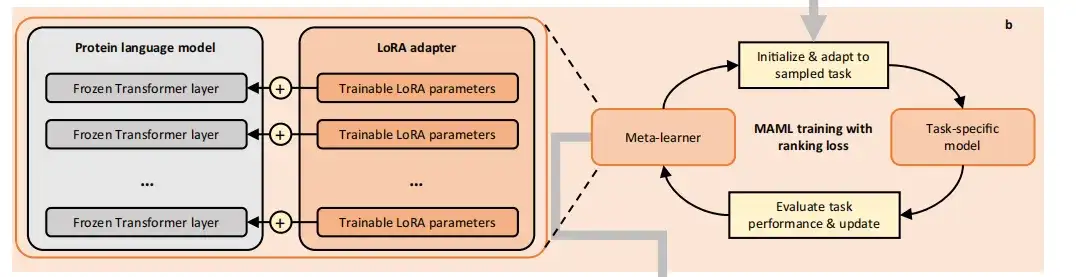
also,The study used a gradient-based meta-learning method called Model-Agnostic Meta-Learning (MAML).Meta-train PLMs on the constructed tasks. MAML is able to find the optimal initial model parameters, even small changes to them will produce significant improvements on the target task. In each iteration cycle, the meta-training process contains two levels of optimization and finally transforms the PLMs into initialized meta-learners.
In the internal optimization, the study uses the current meta-learner to initialize a temporary base learner, which is then updated to a task-specific model by sampling the training data of the task. In the external optimization, the study uses the test loss of the task-specific model on the task to optimize the meta-learner.
In order to avoid catastrophic overfitting due to too little training data,FSFP uses Low Rank Adaptation (LoRA) to inject trainable rank factorization matrices into PLMs.Their original pre-trained parameters are frozen and all model updates are restricted to a small number of trainable parameters.
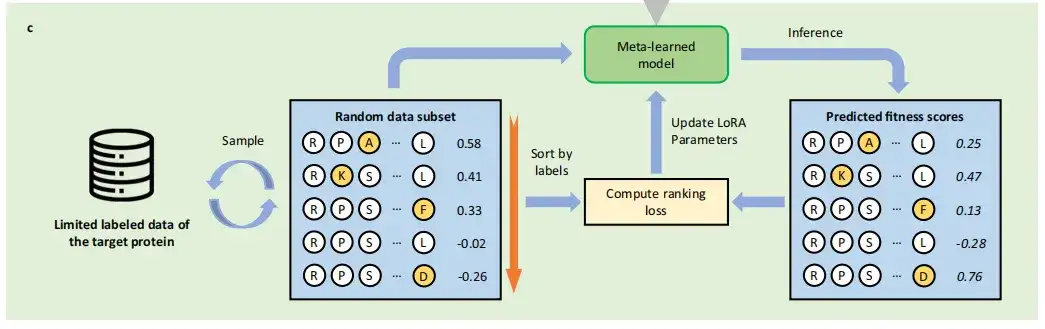
After meta-training, the study can obtain the initialization based on LoRA parameters, and finally transfer the meta-trained PLMs to the target small sample learning task, that is, to learn to predict the mutation effect of the target protein with limited labeled data. Different from the traditional supervised learning method of protein mutation prediction,FSFP regards it as a sorting problem and utilizes LTR technology.
Specifically, FSFP learns to rank mutation fitness by computing the ListMLE loss. In each iteration, the model is trained so that its predictions for one or more sampled data subsets tend toward the ground truth ranking. These training schemes are applied simultaneously to the internal optimization of the transfer learning phase using the target training data and the meta-training phase using the auxiliary task training data.
ProteinGym benchmark based on 87 high-throughput mutation datasets
In order to construct the training tasks required for meta-learning,The method first retrieves existing labeled mutant datasets, retrieves the mutation datasets of the first two proteins closest to the target protein from the largest public collection of DMS datasets, ProteinGym, and uses the MSA-based GEMME pseudo-labeling method to score the mutation information of the target protein to construct a dataset for the third task. These datasets may help predict the effects of mutations on the target protein, and the labeled data for these tasks are randomly divided into training data and test data.
To evaluate the model performance,This study selected the protein mutation dataset (ProteinGym) as the benchmark dataset. The dataset contains approximately 1.5 million missense variants from 87 DMS sequencing experiments. Since the maximum input length of ESM-1v is 1,024, this study truncated proteins with more than 1,024 amino acids and ensured that most of their mutations in the corresponding dataset occurred within the generated interval.
Next, the study randomly selected 20 single-point mutations as the initial training set, and then added 20 single-point mutations to expand the training set size to 40, and then constructed 60, 80, and 100 training sets in the same way. After 5 random data splitting processes,This study can achieve the averaging of model performance on different partitions of a certain training scale.
FSFP is successfully applied to three basic models and has significant advantages in small sample learning tasks
Theoretically, FSFP can be applied to any protein language model based on gradient descent optimization.In order to verify its universality,This study selected three representative PLMs - ESM-1v, ESM-2 and SaPro-t as the basic models for training, and selected the 650M version for evaluation.
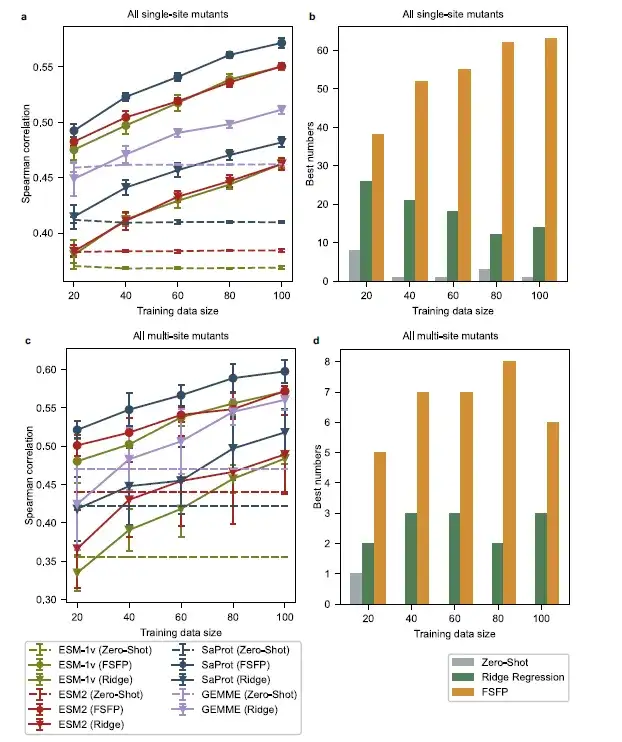
In terms of average performance,PLMs trained with FSFP consistently outperform other baselines across all training data sizes. Among them, SaProt (FSFP) performs best, while ESM-1v (FSFP) and ESM-2 (FSFP) perform comparably. In addition, PLMs trained with FSFP achieve the best Spearman correlation on most datasets in ProteinGym. Compared to zero-shot prediction, FSFP improves the performance of PLMs on single mutants by nearly 0.1 by improving the Spearman correlation of single mutants using only 20 training examples, and this gap becomes even larger when multiple mutants are involved. These improvements continue to increase as the training dataset grows, which is consistent with the ablation experiments of this study.
The model using FSFP achieves significant improvements over GEMME and its enhanced version of ridge regression under all training samples, which indicates that FSFP not only transfers the multiple sequence alignment knowledge from GEMME to PLM, but also successfully combines it with the supervision information from the target training data through multi-task learning.This once again confirms the advantage of FSFP in few-shot learning tasks.
Extrapolation performance evaluation, Spearman correlation evaluation of FSFP trained PLMs is better
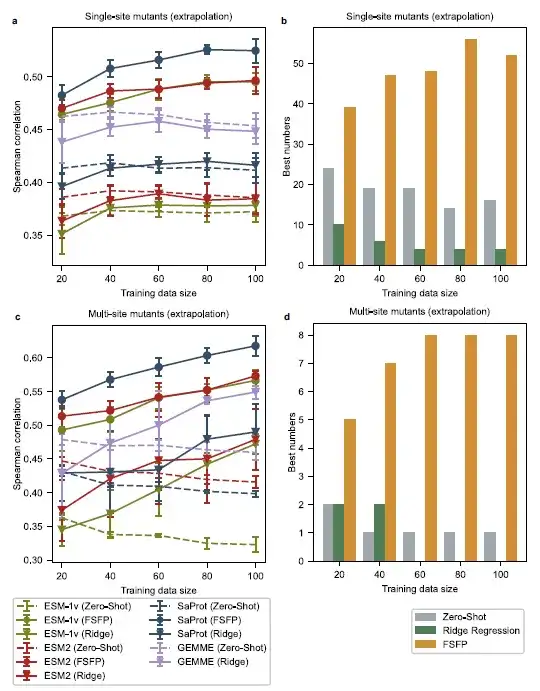
The researchers selected all single-point mutants from each original test set whose mutation sites were different from the training examples, thus obtaining a test set of single-point mutants that were different from the training examples. Then, the researchers selected multi-point mutants whose individual mutations did not overlap with those in the training data, thus obtaining another challenging test set. In this setting, the study found that the zero-shot performance of the base model changed significantly with the size of the training set.
For single-point mutations at different positions, the models enhanced by ridge regression do not perform better than the base models even with 100 training examples. For multiple-point mutations, the ridge regression method is not effective in improving the performance of GEMME and ESM-2 when the training scale is less than 60. In contrast, PLMs trained with FSFP have higher Spearman correlation scores compared to all base models at various training scales. In addition,The best performing models on most datasets are the ones trained with FSFP.
Comprehensive comparison of 4 proteins, FSFP has greater benefits in training with small datasets
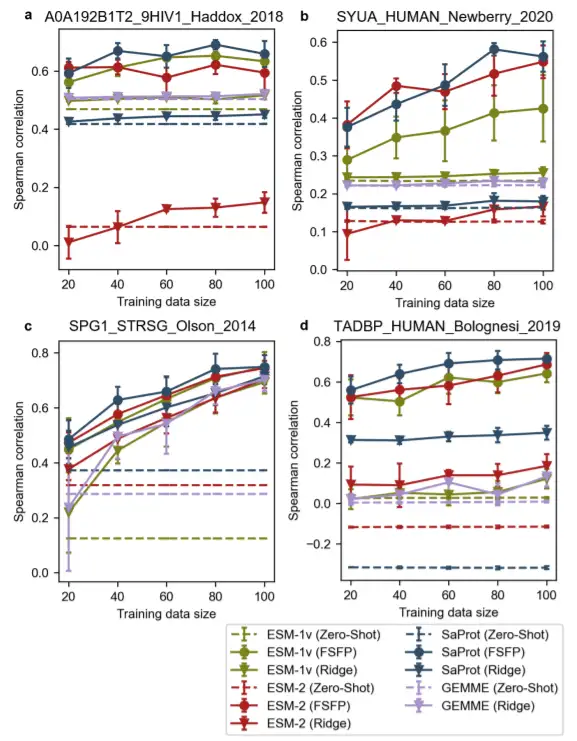
To further demonstrate the applicability and generalization of FSFP,The study also presents comparisons between different methods for four proteins: the envelope protein Env from HIV, the human α-synuclein, protein G (GB1), the human TAR DNA-binding protein 43 (TDP-43). In these cases, one or more unsupervised models performed poorly.
It is noteworthy that for TDP-43, all the spearman correlations for zero-shot predictions are close to zero. Most models augmented with ridge regression, except GB1, also do not achieve significant performance improvements on larger training datasets. In contrast, pre-trained models can achieve considerable gains when trained on small datasets using FSFP.
Using FSFP to design Phi29 DNA polymerase, the positive rate increased by 25%
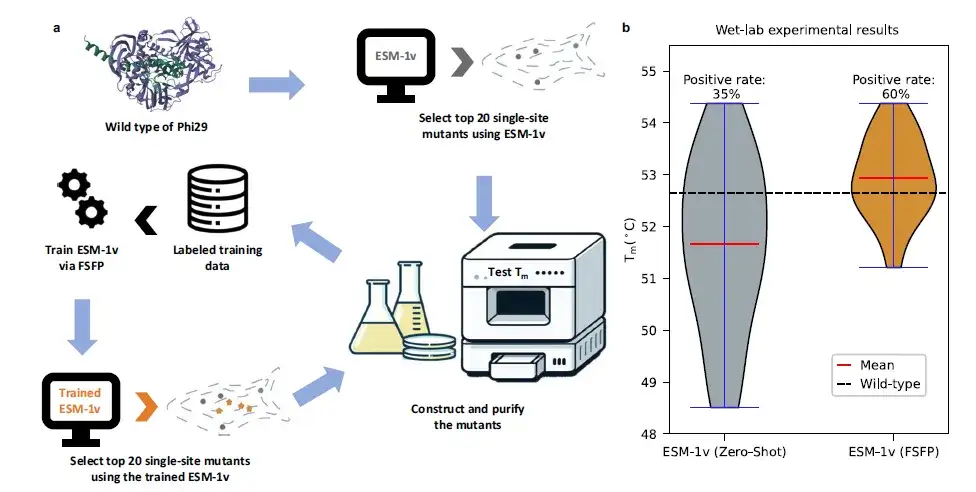
The study also examined a specific case of protein Phi29 modification.Wet test verification was carried out.Based on a limited set of wet experimental data, this study used FSFP to train ESM-1v, used it to find new single-site mutants, and conducted experimental verification. Comparing the top 20 prediction results of ESM-1v before and after FSFP training, the average Tm value increased by more than 1 ℃ and the positive rate increased by 25%.
Specifically, the best mutants (i.e., the mutants with the highest Tm values) found by ESM-1v (FSFP) were also recommended by ESM-1v (zero-shot). However, among the positive mutants predicted by ESM-1v (FSFP), 9 did not appear in the training data, indicating that FSFP can enable PLMs to identify more protein variants.These results confirm the potential of FSFP to accelerate iterative cycles of protein engineering design and testing.This could help develop proteins with enhanced functional characteristics.
A typical representative of AI for Bioengineering, a strong alliance stands at the forefront of the times
Today, when AI and scientific research are closely integrated, we are facing a historic opportunity. Professor Hong Liang believes that although China's biopharmaceutical industry has strong strength, there is still room for improvement in the profit ratio in the international industrial chain. Through AI, we have the opportunity to "change lanes and overtake" and directly use the power of artificial intelligence to promote the development of the industry. It is based on this concept that Professor Hong Liang and researcher Tan Pan have launched endless explorations in the field of AI for Bioengineering.
Dr. Tan Pan focuses on molecular biophysics, artificial intelligence functional protein design, and drug molecule design.Published 15 SCI papers in journals such as Nature Communications, PRL, Journal of Cheminformatics, PCCP, etc. Developed a variety of AI-assisted protein design and modification algorithms. Combining Professor Hong Liang's professional technology with Dr. Tan Pan's AI algorithm, the collaborative research between the two parties has achieved many results.
Over the years, both parties have focused on innovative research on general artificial intelligence in the field of protein engineering and have successfully developed the pro series of protein engineering general artificial intelligence. Similar to the way ChatGPT understands human language, the pro series uses large models to understand the amino acid arrangement of proteins in nature and designs protein products with superior performance. Among them, there are also two major milestone products in industrial application:
* Extremely alkali-resistant single domain antibody:The world's first large-scale model-designed protein product, jointly developed with Jinsai Pharmaceuticals, has achieved 5,000 liters of industrial production, providing a new solution for the purification of biological macromolecules.
* Glycosyltransferase:Cooperating with Hanhai New Enzyme, we developed the enzyme for producing EPS-G7, the core material for pancreatitis screening, breaking the long-term foreign monopoly and significantly reducing costs.
These two cases mark the world's first and second large-scale protein products that have been designed and successfully scaled up for industrialization. Based on his deep accumulation in the field of AI protein design, Professor Hong Liang founded Shanghai Tianwu Technology Co., Ltd. in 2021. In just three years, the company has not only completed multiple protein design projects, but also obtained tens of millions of yuan in Pre-A rounds of financing from well-known institutions such as Glory Ventures and GSR Ventures.
At present, the company's services have covered multiple fields such as innovative drugs, in vitro diagnostics, synthetic biology, etc., and it is actively seeking cooperation with more scientific research institutes and enterprises, and is committed to setting a national and even global benchmark in the field of protein engineering.
In the fiercely competitive field of protein engineering, Professor Hong Liang’s vision is clear:We must not only become a domestic leader, but also a global leader.In the future scientific research journey, Professor Hong Liang and his team are committed to expanding in-depth cooperation with global scientific research institutions and enterprises, constantly exploring the infinite possibilities of protein design, striving to achieve technological breakthroughs and application innovation in this field, setting a benchmark domestically and demonstrating excellence internationally.
Finally, I recommend an online academic sharing activity. Interested friends can scan the QR code to participate!