Command Palette
Search for a command to run...
Directly Attacking Three Major Solid Tumors! Shanghai Jiaotong University Team Released a Deep Learning System to Improve the Accuracy of Cancer Survival Prediction

A report released by the World Health Organization in 2022 pointed out thatNon-communicable diseases (NCDs) such as cancer have surpassed infectious diseases to become the world's number one killer.The latest data released by the National Cancer Center of China show that there will be approximately 4.8247 million new cancer cases and 2.5742 million new cancer deaths in China in 2022.
For a long time, people have been afraid of cancer. But as a chronic disease, 1/3 of cancer is preventable, 1/3 can be cured through early detection, early diagnosis, and early treatment, and 1/3 is incurable, but can be controlled through proper treatment, with a better quality of life and longer survival. Prevention is mainly through improving one's own immunity, regular physical examinations, and paying attention to personal health. After cancer is diagnosed, prognosis analysis is very important.
Cancer prognosis refers to the prediction of the possible course and outcome of cancer patients. Prognostic analysis helps improve the survival chances of cancer patients.In the past, researchers have characterized the tumor microenvironment (TME) from the perspective of spatial gene expression based on spatial transcriptomics (ST) technology to distinguish different prognostic subgroups of cancer patients. However, the high cost and long experimental cycle of ST have hindered its application in large-scale cancer patient cohorts for survival prediction. In contrast, histological images are cost-effective, easy to obtain in clinical settings, and can provide rich information about tumor morphology. They are a better alternative to molecular-level TME analysis and can achieve more accurate cancer prognosis.
Recently, the research group of Yu Zhangsheng (School of Life Sciences and Technology/Clinical Research Center of School of Medicine), the research group of Wang Yuguang (School of Natural Sciences/School of Mathematical Sciences) of the Shanghai National Center for Applied Mathematics (Shanghai Jiao Tong University Branch) and their collaborators published a paper titled "Harnessing TME depicted by histological images to improve cancer prognosis through a deep learning system" in Cell Reports Medicine.This study developed a deep learning system that can predict tumor microenvironment information for cancer patients without spatial transcriptome data through histopathology images, thereby achieving accurate cancer prognosis.
Research highlights:
- Predicting TME information from histopathological images for cancer patients without ST data
- TME characterized by IGI-DL improves the accuracy of cancer survival prediction
- Significantly expanded the use of gene spatial expression information in large public databases of biomedical pathology images

Paper address:
https://www.cell.com/cell-reports-medicine/fulltext/S2666-3791(24)00205-2
Follow the official account and reply "tumor microenvironment" to get the complete PDF
Dataset: Evaluating tissue samples from 3 solid tumor types
This study used three different datasets to evaluate the performance of the model on tissue samples of three different solid tumor types: colorectal cancer (CRC), breast cancer, and cutaneous squamous cell carcinoma (cSCC).
For colorectal cancer,The researchers used 41,492 points from 10 ST datasets of 10 CRC patients from Ruijin Hospital affiliated to Shanghai Jiao Tong University School of Medicine, which were sequenced by 10× Visium as a leave-one-patient-out validation set, as shown in the table below.

For breast cancer,The researchers used 34,678 spots from 92 tissue samples from 27 patients, which were sequenced using traditional ST technology, as a leave-one-patient validation set, as shown in the table below.

For squamous cell carcinoma of the skin,The researchers used 4,353 spots from 12 tissue samples from four patients that were sequenced using traditional ST technology as a leave-one-patient validation set, as shown in the table below.

Model architecture: New deep learning system improves cancer prognosis
In this study, researchers developed a deep learning system that can improve cancer prognosis using the TME depicted in histological images.
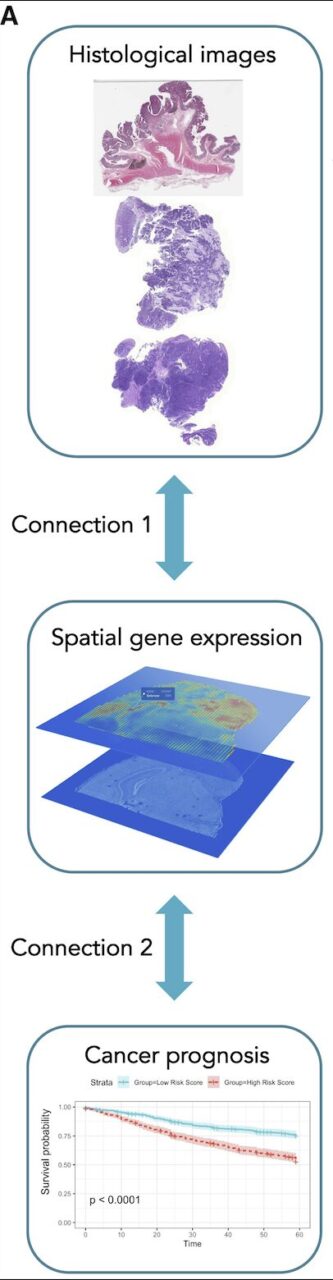
The system consists of two parts:
The first part (Connection 1 in the figure above) is a model based on integrated graph and image deep learning (IGI-DL), which uses convolutional neural networks and graph neural networks to project H&E stained histological images into gene expression space.
In the second part (Connection 2 in the figure above), the researchers used the super-patch graph and IGI-DL predicted spatial gene expression as node features in the colorectal cancer cohort and breast cancer cohort in the Cancer Genome Atlas (TCGA) dataset to predict prognosis, and then verified it in the external test set MCO-CRC (Molecular and Cellular Oncology colorectal cancer).
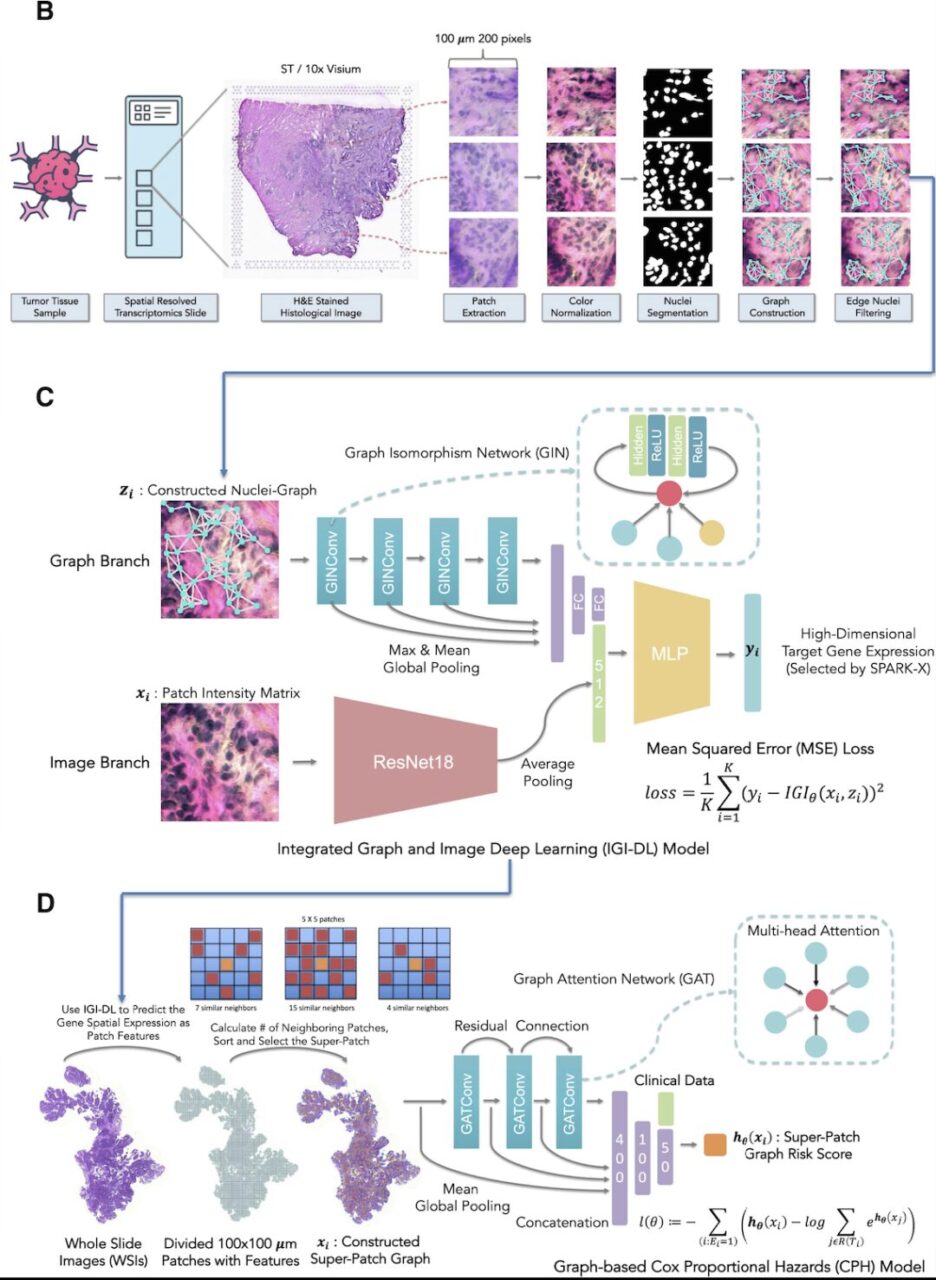
Specifically, the construction of the system includes three steps: H&E staining histological image preprocessing, spatial gene expression prediction model and spatial gene expression super-patch graph survival model based on the prediction.
- H&E-stained histological image preprocessing:First, each H&E-stained histological image was segmented into multiple non-overlapping patches of 200 × 200 pixels with a resolution of 0.5 μm/pixel according to the coordinates of each point;
- Spatial gene expression prediction model:For each patch, the researchers built a Nuclei-Graph, in which each nucleus segmented by Hover-Net24 was represented as a node, and the distance between each pair of nuclei determined whether there was an edge connection. Based on the architecture shown in Figure C above, the researchers used the IGI-DL model to predict the target gene expression at each point in the histological image.
- Super-patch graph survival model based on predicted spatial gene expression:To further predict prognosis based on TME delineated by spatial gene expression, the researchers constructed a super-patch graph from H&E-stained whole-slide imaging (WSI) of each patient with cancer, and then constructed a graph-based survival prediction model using the constructed super-patch graph and clinical characteristics as input.
Research results: IGI-DL model performs well overall
Overall, the IGI-DL model constructed in this study integrates the advantages of convolutional neural networks and graph neural networks, making full use of the pixel intensity and structural features in histopathological images to achieve more accurate prediction of gene spatial expression levels.The model performed well in three types of solid tumors: colorectal cancer, breast cancer, and cutaneous squamous cell carcinoma, with an average correlation coefficient improvement of 0.171 compared with five existing methods.

For colorectal cancer, the researchers compared the Pearson correlation of 179 genes predicted by IGI-DL with five SOTA models.IGI-DL achieved an average Pearson correlation of 0.343 in the 10 retained patients, significantly outperforming the other models with an average increase of 0.233,As shown in the picture above.

For breast cancer, the researchers compared the Pearson correlation of 187 genes predicted by IGI-DL with the previous model, and IGI-DL achieved an average correlation of 0.231 in the 27 retained patients. As shown in the figure above,The IGI-DL model outperforms all SOTA models with an average improvement of 0.142.

For cutaneous squamous cell carcinoma, the researchers compared the Pearson correlation of 487 genes predicted by IGI-DL with previous models. IGI-DL achieved an average correlation of 0.198 in the four retained patients, which was the best performance among all models.The average performance of other SOTA models is improved by 0.131.As shown in the picture above.
In terms of cross-platform and cross-cancer performance, as in the above experiments, the best SOTA model is not fixed for internal validation and external test sets of different cancer types.However, the performance of the IGI-DL model is always better than other models, with an average improvement of 0.171, showing good cross-platform generalization ability.
Furthermore, the researchers investigated the cross-cancer prediction performance of IGI-DL, and the model trained on colorectal cancer performed well on the internal validation and external test sets of cutaneous squamous cell carcinoma, with average correlations of 0.204 and 0.143, respectively. However, most cross-cancer prediction performances were lower than when training and testing a single cancer type.This result suggests that spatial gene expression in tumor regions is somewhat cancer-specific, and cross-cancer prediction is inherently difficult.

Regarding prognostic prediction performance, in the Cancer Genome Atlas Breast Cancer (TCGA-BRCA) cohort, the super-patch graph survival model based on spatial gene expression as node features can achieve an average consistency index (C-index) of 0.747 in 5-fold cross-validation; in the Cancer Genome Atlas Colorectal Cancer (TCGA-CRC) cohort, the survival model has a C-index of 0.725 in 5-fold cross-validation, which is better than other prognostic models, as shown in the figure above.
The survival prognosis model also maintains an accuracy advantage for the prognosis prediction of early-stage patients (stage I and II), and the predicted risk score can be used as an independent prognostic indicator for patients of all stages and early-stage patients. In the external test set MCO-CRC containing data from more than a thousand patients, the survival prognosis model maintains a stable advantage and has generalization ability.
Breast cancer and pancreatic cancer first: Leveraging AI to improve prognosis
During the diagnosis and treatment of cancer, cancer prognosis analysis can effectively avoid overtreatment and waste of medical resources, and provide a scientific basis for medical staff and their families to make medical decisions. It has become a hot topic in cancer research in recent years.
To improve breast cancer outcomes, in 2020, Salesforce researchers collaborated with clinicians at the Lawrence J. Ellison Research Institute at the University of Southern California.Launched the machine learning system ReceptorNet,Its algorithm can predict hormone receptor status from low-cost and easily accessible tissue images—an important biomarker for clinicians when deciding the appropriate treatment path for breast cancer patients. The system has an accuracy rate of 92%.

In February 2024, researchers from the University of Kentucky, Macau University of Science and Technology, University of Macau, and the First Affiliated Hospital of Guangzhou Medical University used a neural network model toA precise prognostic scoring system, MIRS (metastasis and immunogenomic risk score), was established.The tumor metastasis and immune genomic risk scoring provides a predictive tool that is almost universally applicable to breast cancer patients and provides a new direction for treatment options for the breast cancer population. (Click here for detailed report: Aiming at the world's most common cancer, Chinese scholars established the breast cancer prognostic scoring system MIRS)
In addition, pancreatic cancer is one of the common malignant tumors of the digestive tract, and the five-year survival rate after diagnosis is no more than 10%. A key link in improving the survival rate of patients is to accurately predict the prognostic risk of patients in order to design targeted treatment plans. Histopathology is a routine examination in the oncology department. It can analyze the characteristics of tumors at the microscopic level and is an important method for assessing the risk of tumor progression. However, due to the large size of the slices and the complex tissue composition, the evaluation results are easily affected by subjective factors.

In 2023, a research team from the Nanjing University of Information Science and Technology and the Institute of Smart Healthcare, School of Artificial Intelligence,Published a research paper titled "Multi-tissue segmentation model for pancreatic cancer whole-slice images based on multi-task and attention"The tissue segmentation of 8 categories of pancreatic cancer pathological sections was studied. By introducing an attention mechanism and designing a hierarchical and shared multi-task structure, the model performance was significantly improved by using related auxiliary tasks.
The model proposed in this study was trained and tested on the dataset of Shanghai Changhai Hospital and externally validated on the TCGA public dataset. The F1 scores on the internal test set were higher than 0.97, and the F1 scores on the external validation set were higher than 0.92. The generalization performance was significantly better than the baseline method.
It is worth emphasizing that AI cannot replace pathology experts, but as an auxiliary diagnosis technology, it brings more convenience to pathology diagnosis and further improves the work efficiency of pathologists. From a long-term perspective, AI still has a lot of room for development in digital biomarker detection, medical image analysis, and disease course prediction.
References:
1.https://news.un.org/zh/story/2018/09/1017602
2.https://mp.weixin.qq.com/s/VE68FKL6kwpO1IFsbR-LVA
3.https://ins.sjtu.edu.cn/articles/286
4.https://www.cdstm.cn/theme/khsj/khzx/khcb/202012/t20201214_1039028.html








